CLPC.1 (Potri.012G105900) [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | CLPC.1 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.012G105900.3 pacid=42783939 polypeptide=Potri.012G105900.3.p locus=Potri.012G105900 ID=Potri.012G105900.3.v4.1 annot-version=v4.1
ATGGCTAGGCTTCTAGTTCATTCTGCTAATATACCTGCTGTAGCTCCTTGTCCAAGGCATTGTCAATATGAAGAATCTAAAAAATCAAGGGCCTCCTCTG
CCAAAATGATGTGTAGCTTACCATCGCGGGGTTTGGTGATAAGTGGATACTCAGGATTGCGTAGCGCAAACTGTTTAGATACACTGTTGAGACATGGACA
CTCTTTCCATTCTAAGGTTGCAATTACAATCTCTCCTAGGCAGCAAAAGGCTAAGCGATTTGTACCTAGAGCTATGTTTGAACGTTTTACAGAAAAAGCA
ATCAAAGTAATTATGCTTGCTCAAGAGGAAGCAAGACGGCTTGGTCATAATTTTGTCGGAACAGAACAGATCCTGTTGGGGCTTATTGGTGAAGGAACTG
GAATTGCTGCAAAGGTTCTCAAATCTATGGGAATCAATCTTAAAGATGCACGTGTAGAGGTGGAGAAGATTATTGGCAGGGGTAGTGGATTTGTTGCTGT
GGAAATTCCGTTCACTCCTCGTGCCAAGCGTGTTTTGGAACTATCATTGGAAGAGGCCCGCCAACTTGGTCATAATTATATTGGATCCGAGCACTTGCTT
CTCGGGCTACTTCGTGAAGGTGAAGGTGTAGCAGCCCGTGTTCTCGAGAATTTAGGTGCTGATCCTAGTAACATACGTACACAGGTCATTCGTATGGTGG
GTGAGAGCACAGAAAACCTGGCTGGTTCCACTGTTGGACCTGGAAGCAGTAACAACAAGATGCCTACACTTGAGGAATATGGAACTAATTTGACTAAATT
AGCAGAGGAGGGTAAATTGGATCCTGTTGTCGGAAGGCAGCCACAAATAGAACGTGTTGTTCAAATCCTGGGCCGAAGAACTAAGAATAACCCTTGTCTC
ATTGGAGAACCTGGGGTTGGGAAAACTGCAATTGCTGAAGGTCTTGCTCAGAGGATAGCAAGCGGTGATGTTCCTGAGACAATTGAGGGAAAGAAGGTTA
TCACCCTTGACATGGGCCTTCTTGTTGCTGGAACTAAATACCGTGGAGAGTTTGAGGAAAGATTGAAGAAGCTAATGGAGGAAATCAAACAGAGTGATGA
AATAATTCTCTTTATTGATGAGGTGCATACCCTGATAGGGGCAGGAGCAGCAGAAGGGGCTATTGATGCTGCAAATATTTTGAAACCAGCTCTTGCTAGA
GGTGAATTGCAGTGTATTGGAGCTACAACCCTAGATGAGTACAGAAAGCACATTGAGAAGGACCCTGCTTTGGAAAGACGTTTCCAGCCAGTCAAAGTAC
CTGAGCCATCTGTGGATGAAACAATACAGATTTTGAAAGGACTTCGTGAGCGATATGAGATCCACCACAAACTCCGTTACACAGATGAGGCCTTAGTAGC
TGCTGCTCAGTTGTCATACCAGTACATCAGTGACCGTTTTCTGCCTGATAAAGCAATTGATCTGATCGATGAAGCTGGTTCTCGGGTTCGCCTTCGGCAT
GCACAGGTCCCTGAGGAAGCAAGAGAGCTTGAGAAAGAGGTCAGACAAATCACAAAGGAAAAGGATGAAGCTGTCCGCGGCCAAGACTTCGAAAAGGCTG
GGGAATTGCGAGACAGAGAAATGGACCTTAGAGCACAGATCGCAGCTATTGTGGAGAAAGGCAAGGAAATGAGCAAAGCTGAGACAGAGGCTGGGGATGT
AGGTCCCACTGTCACTGAATCAGATATTCAGCACATTGTCTCTTCTTGGACTGGCATACCTGTAGAGAAAGTATCCACGGATGAATCTGATCGTCTCCTA
AAGATGGAAGATACCCTTCACAAGAGAGTTATTGGTCAGGATGAAGCTGTCAAGGCAATCAGCCGTGCCATCCGCCGAGCTCGTGTTGGGCTTAAGAACC
CAAATCGACCAATTGCTAGCTTCATCTTTTCTGGTCCAACTGGTGTCGGGAAGTCAGAACTGGCAAAAGCACTAGCTGCCTACTACTTTGGCTCAGAAGA
AGCCATGATCCGACTTGATATGAGCGAGTACATGGAAAGACACACTGTTGCCAAGCTAATTGGTTCACCTCCTGGTTATGTTGGTTACACTGAAGGTGGT
CAGTTGACAGAGGCTGTTCGGCGTCGTCCTTACACTGTGGTACTCTTTGATGAGATTGAGAAGGCCCATCCTGATGTCTTTAATATTATGCTTCAAATTC
TTGAAGATGGAAGGTTGACAGACAGCAAGGGCAGGACAGTGGACTTCAAAAATACCCTCCTGATAATGACTTCAAATGTTGGTAGCAGTGTTATTGAGAA
GGGAGGACGCAAGATGGGATTTGATCTTGATTATGATGAGAAGGATAGCAGTTACAACCGAATTAAGAGCCTGGTGACAGAGGAGTTAAAGCAGTATTTT
AGGCCAGAATTCTTGAACAGATTGGATGAGATGATTGTTTTCCGTCAACTTTCCAAGCTGGAGGTCAAGGATATTGCTGATATAATGTTGAAGGAGGTCT
TTGAGAGGTTGAAGGCCAAGGAAATAGAACTTCAAGTGACTGAGAGATTTAGAGATAGGGTGGTCGATGAAGGGTATAACCCATCCTATGGTGCAAGGCC
TTTGAGAAGAGCTATAATGAGGCTCCTTGAGGACAGCATGGCTGAGAAAATGCTTTCAGCAGAGATCAAAGAAGGTGATTCTGTCATTATAGATGTTGAT
TCTGATGGAAATGTTGTTGTGCTCAATGGTCAAAGTGGTGGTGCTCCCGATGCCTTGCCCGACATGTTGAACGTTGTGTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.012G105900.3 pacid=42783939 polypeptide=Potri.012G105900.3.p locus=Potri.012G105900 ID=Potri.012G105900.3.v4.1 annot-version=v4.1
MARLLVHSANIPAVAPCPRHCQYEESKKSRASSAKMMCSLPSRGLVISGYSGLRSANCLDTLLRHGHSFHSKVAITISPRQQKAKRFVPRAMFERFTEKA
IKVIMLAQEEARRLGHNFVGTEQILLGLIGEGTGIAAKVLKSMGINLKDARVEVEKIIGRGSGFVAVEIPFTPRAKRVLELSLEEARQLGHNYIGSEHLL
LGLLREGEGVAARVLENLGADPSNIRTQVIRMVGESTENLAGSTVGPGSSNNKMPTLEEYGTNLTKLAEEGKLDPVVGRQPQIERVVQILGRRTKNNPCL
IGEPGVGKTAIAEGLAQRIASGDVPETIEGKKVITLDMGLLVAGTKYRGEFEERLKKLMEEIKQSDEIILFIDEVHTLIGAGAAEGAIDAANILKPALAR
GELQCIGATTLDEYRKHIEKDPALERRFQPVKVPEPSVDETIQILKGLRERYEIHHKLRYTDEALVAAAQLSYQYISDRFLPDKAIDLIDEAGSRVRLRH
AQVPEEARELEKEVRQITKEKDEAVRGQDFEKAGELRDREMDLRAQIAAIVEKGKEMSKAETEAGDVGPTVTESDIQHIVSSWTGIPVEKVSTDESDRLL
KMEDTLHKRVIGQDEAVKAISRAIRRARVGLKNPNRPIASFIFSGPTGVGKSELAKALAAYYFGSEEAMIRLDMSEYMERHTVAKLIGSPPGYVGYTEGG
QLTEAVRRRPYTVVLFDEIEKAHPDVFNIMLQILEDGRLTDSKGRTVDFKNTLLIMTSNVGSSVIEKGGRKMGFDLDYDEKDSSYNRIKSLVTEELKQYF
RPEFLNRLDEMIVFRQLSKLEVKDIADIMLKEVFERLKAKEIELQVTERFRDRVVDEGYNPSYGARPLRRAIMRLLEDSMAEKMLSAEIKEGDSVIIDVD
SDGNVVVLNGQSGGAPDALPDMLNVV
|
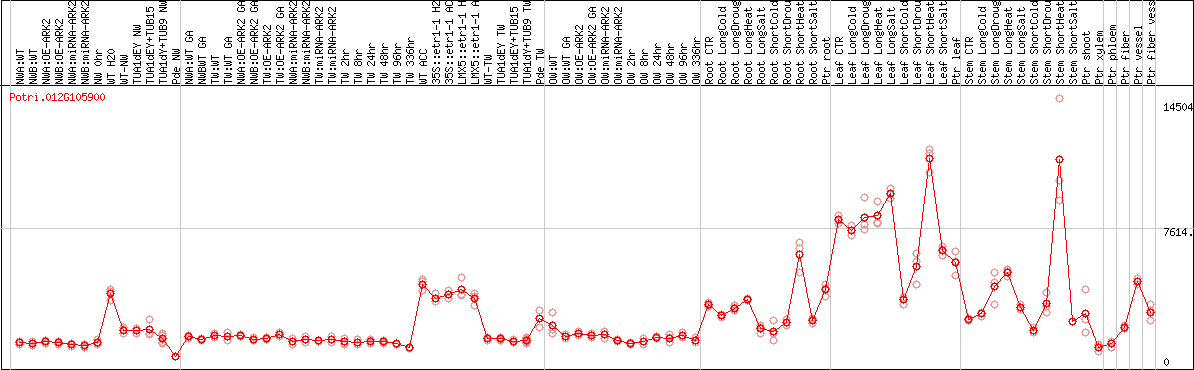
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.012G105900 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
