External link
Symbol
DREB36,DREB1.2
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G46768 160 / 3e-51
AP2_ERF
RAP2.1
related to AP2 1 (.1)
AT5G67190 158 / 5e-50
AP2_ERF
DEAR2
DREB and EAR motif protein 2 (.1)
AT3G50260 153 / 2e-48
AP2_ERF
DEAR1, CEJ1, ATERF#011
DREB AND EAR MOTIF PROTEIN 1, cooperatively regulated by ethylene and jasmonate 1 (.1)
AT2G23340 151 / 4e-47
AP2_ERF
DEAR3
DREB and EAR motif protein 3 (.1)
AT4G36900 149 / 2e-46
AP2_ERF
DEAR4, RAP2.10
DREB AND EAR MOTIF PROTEIN 4, related to AP2 10 (.1)
AT4G06746 146 / 1e-45
AP2_ERF
DEAR5, RAP2.9
DREB AND EAR MOTIF PROTEIN 5, related to AP2 9 (.1)
AT4G16750 98 / 2e-26
AP2_ERF
Integrase-type DNA-binding superfamily protein (.1)
AT1G71450 97 / 4e-26
AP2_ERF
Integrase-type DNA-binding superfamily protein (.1)
AT1G21910 95 / 1e-24
AP2_ERF
DREB26
dehydration response element-binding protein 26, Integrase-type DNA-binding superfamily protein (.1)
AT2G35700 94 / 1e-24
AP2_ERF
ATERF38
ERF family protein 38 (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10018727
196 / 1e-64
AT5G67190 162 / 1e-51
DREB and EAR motif protein 2 (.1)
Lus10010043
182 / 1e-59
AT2G23340 170 / 1e-54
DREB and EAR motif protein 3 (.1)
Lus10009373
155 / 4e-49
AT5G67190 164 / 3e-52
DREB and EAR motif protein 2 (.1)
Lus10009798
97 / 6e-26
AT1G19210 132 / 1e-39
Integrase-type DNA-binding superfamily protein (.1)
Lus10038082
97 / 3e-25
AT5G21960 155 / 3e-47
Integrase-type DNA-binding superfamily protein (.1)
Lus10043240
96 / 7e-25
AT5G11590 176 / 1e-54
TINY2, Integrase-type DNA-binding superfamily protein (.1)
Lus10001601
95 / 1e-24
AT5G11590 159 / 1e-48
TINY2, Integrase-type DNA-binding superfamily protein (.1)
Lus10033420
94 / 2e-24
AT1G19210 181 / 4e-58
Integrase-type DNA-binding superfamily protein (.1)
Lus10002801
94 / 3e-24
AT5G11590 209 / 4e-68
TINY2, Integrase-type DNA-binding superfamily protein (.1)
Lus10019665
92 / 2e-23
AT4G39780 209 / 1e-67
Integrase-type DNA-binding superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0081
MBD-like
PF00847
AP2
AP2 domain
Representative CDS sequence
>Potri.014G025200.3 pacid=42764403 polypeptide=Potri.014G025200.3.p locus=Potri.014G025200 ID=Potri.014G025200.3.v4.1 annot-version=v4.1
ATGGAAGGAGGGTGCTGTACATCGTCAACAAGTACTCCAAGTTCAACTTCAACAGCAACCACAGAGAAACGGAAGCATAGCAGGCAACAAAACCAAGAGA
AGCCATACAGAGGGATAAGGATGAGGAAGTGGGGTAAGTGGGTAGCAGAAATTAGAGAACCAAACAAGAGGTCTAGGATTTGGCTTGGGTCCTACTCAAC
ACCAATAGCTGCAGCTCGTGCATACGACACGGCTGTTTTCTATCTGCGAGGCCCTTCTGCTAGGCTTAATTTCCCTGATTTGATATACCAAGAAGATGAG
CTGCGAGATGTGTCTGCTGCTTCTATACGCAAGAAAGCCACTGAAGTTGGGGCTAAAGTGGATGCCTTACAAACCGCTGTCCATGCATCACCAGAAGATA
ACTCACGAGTGTTACTTTCAGAGAAACCTGATTTGAACAAGTTCCCAGAAAATTCTGATGAAGAGTGA
AA sequence
>Potri.014G025200.3 pacid=42764403 polypeptide=Potri.014G025200.3.p locus=Potri.014G025200 ID=Potri.014G025200.3.v4.1 annot-version=v4.1
MEGGCCTSSTSTPSSTSTATTEKRKHSRQQNQEKPYRGIRMRKWGKWVAEIREPNKRSRIWLGSYSTPIAAARAYDTAVFYLRGPSARLNFPDLIYQEDE
LRDVSAASIRKKATEVGAKVDALQTAVHASPEDNSRVLLSEKPDLNKFPENSDEE
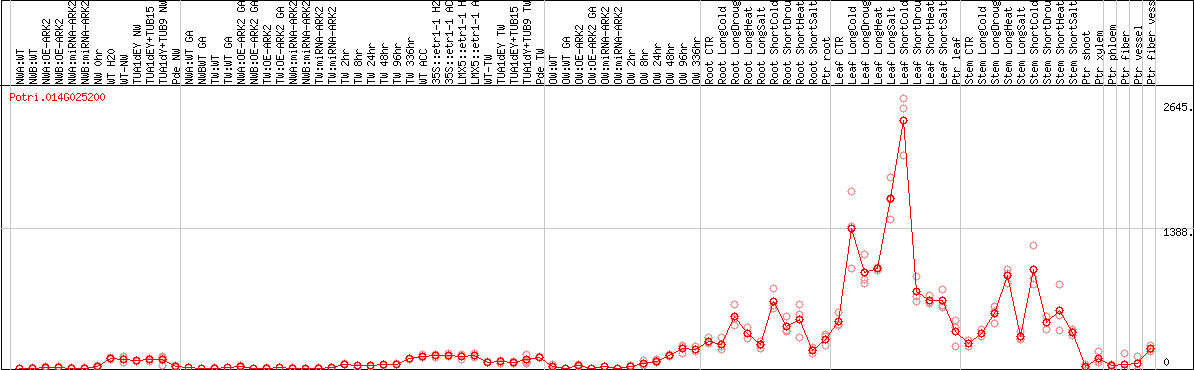
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.014G025200 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.