Potri.017G015200 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.017G015200.1 pacid=42813172 polypeptide=Potri.017G015200.1.p locus=Potri.017G015200 ID=Potri.017G015200.1.v4.1 annot-version=v4.1
ATGGCTCTGGAGTTGATCGGAGGATCAATTCTCTCTGCTCTCGTTGAGGTACTGGTCGACAGGTTGGCCTCTCGTGAGGTTCTGGACTCCTTCAAAAGCC
ACAAACTCGATGATGGTCTTCTGGAAAAGCTGAATGAAACACTGAATACTGTCAACGGACTACTCGACGATGCGGAGGAGAAGCAGATTACAAAGCGTGC
TGTGAAGAACTGGCTTAATGATGTTAAACACGCTGTGTATGAAGCTGAGGACATATTGGATGAGATTAATTATGAATATCTACGATCCAAGGACATTGAT
ACCCCTCGACCTGACAGCAATTGGGCAAGGAATCTGGTTCCATTTCTAAATCCAGCTAATAGAAGGATGAAAGAGATTGGAGCAGAGTTACAAAAGATCT
ATGAGAAGCTTGAACGTTTACTTAAACACAAGGGTGATCTGCGCCACATAGAAGGTACTGGTGGATGGAGACCATTGTCAGAGAAAACAACTCCACTGGT
TAATGAACCTGATGTATATGGAAGGGACGCTGATAAGGAAGCCATAATGGAGCACTTGCTGACACAACACAAGACAGATGGCCCAAATTTAGTTGTGGTT
CCTATTGTGGGCATGGGCGGGATTGGTAAAACCACTCTTGCTCAGCTTATCTACAACGACGAAAAGGTAGATCAGTTCTTCCAACTCAAGGCCTGGGTCT
GGGCTTCTCAACAATTTGACGTCACCAGGATTATTGAGGATATTATTAAGAAGATCAAAGCAAGGACTTGCCCCACCAAAGAACCAGATGAATCCAAAGA
ACCAAATGAATCTCTAATGGAGGCAGTGAAAGGGAAAAAACTTCTACTTGTTCTAGATGACGCGTGGAACATCGAGTACAATGAATGGGATAAATTACTG
CTACCTTTGCGGTATGTGGAGCACGGAAGTAAGATTGTCGTGACAACACGGGAAGAAGATGTAGCAAAAGTCACGCAGACTGTCATCCCCTCTCACCGCT
TGAATGTAATAAGCGATGAAGATTGCTGGAAGTTGTTTGCAAGGGATGCATTTAGTGGTGTAAATTCTGGAGCAGTCTCACACTTGGAAGAATTTGGCAG
AGTAATAGTGAGAAAGTGCAAAGGTTTACCGCTGGCAGCGAAAACTCTTGGGGGTCTTTTGCACTCCGTAGGAGATGTCAAGCAATGGGAGAAGATATCC
AATAGCAGTATGTGGGGTTCGTCGAATGAAAACATCCCTCCAGCTCTAACATTGAGCTATTATTATCTCCCTTCACACCTGAAACGTTGTTTTGCTTATT
GTGCAATATTTCCCAAGGATTATGTATTCAAGAAGGATAGATTAATCACTGAATGGATGGCACATGGTTTTCTAGTCCAGCCCAGAGGCGTTGAGGAGAT
GGAAGATATAGGTGAAAAATACTTTAATGATCTTGTCTCCAGGTCACTTTTCCAGCAATCCACTGGTGATTCTTTTTTCAGCATGCATGACCTCATAAGT
GATTTAGCTGAATACGTGTCAGGAGAATTTTGCTTCAAGCTTGGTATTAATGAATCGGGCTCTGGGTTGGAGAGTGAACATTCATGCTCGCTTCCAGAAA
GGACTCGTTATTTATCAATCACTTCAGCGGCAGCATATGGCGGAGGACTTCGGATTTTTCGTAGCATTCATGGAGTCCAACATTTGCGCGCCTTGTTTCC
ACTGAAGTTTTTTGTGGAAGTTGATATTGAGGCACTGAATGACATTTTGCCAAATCTTAAGCGCTTGCGAATGCTATCTTTATGTCATCCAAAAGATATT
TCTTCTCAGTTGCTCAATTCCATTGGCAACTTAAAACATTTGCGGCACTTGGACCTTTCTCAGACAGTATTTAAACGGTTGCCCGAAAGTGTGTGCACCT
TGTACTATTTGCAAAGCTTATTGTTAAAGGAGTGCCGACTTCTCATGGAGTTGCCATCCAACCTTTCCAACTTGGTCGACTTACAACATCTTGATATTGA
AGGGACAAATTTGAAAGAGATGCCACCAAAAATGGGGAAACTCACGAAGCTTCGAATTTTGGAATCTTACATCGTGGGAAAAGATAGCGGGTCTAGCATG
AAAGAGTTGGGGAAGCTCTCACACATAAGGAAAAAACTTTCTATTCGGAATCTCAGAGATGTTGCAAATGCTCAAGATGCTTTGGATGCCAATTTGAAGG
GTAAGAAGAAGATTGAGGAGTTGGGGTTGACGTGGGATGGCAGTACGGATGACACACCACACGAGAGAGATGTACTTGAGAAATTGGAGCCTTCTGAAGA
TGTGAAAGAGCTTGCCATTATTGGTTACGGGGGTACAACGTTTCCAGGCTGGCTTGGAAACTCTTCTTTCTCAAATATGGTAACGTTGCTTCTTTCTGGA
TGTACGAACTGCATCCTCTTACCACCATTGGGGCAGCTGCCATCTCTAGAAGAGCTCGAAATTGAAGGATTTGATGAAGTCGTGGCAGTTGGTTCTGAGT
TCTATGGAAGTGACCCTCCAATGGAGAAGCCATTTAAATCCCTCATAACATTAAAGTTTGAGGGGATGAAGAAATGGCAGGAATGGAATACAGATGTAGC
TGGTGCTTTCCCTCATCTTGAAAATCTCTTGATAGCAGGTTGTCCAGAGTTAACAAATGGCTTGCCTAATCACCTTCCTTCTTTATTGATACTTGAGATT
CGAGCGTGTCCGCAGCTTGTGGTTTCAATTCCAGAGGCTCCCCTGCTCAGAGAAATCAATGTATCTGAGGGAGATGAAAGTCGTATAACTGGTCGTACGT
CGTCCTATTATTGCTACTACTTACCAGATCAACCACCCAAGTGCCTTCATTTTAGACGAGATCCCCAGCTAACGGGAATGGAGCAAATGAGTCATTTGGA
TCCCCGTCTTTTCACTGATATCAAAATCGAGGAATGTAGTTCATTTAAGTGCTGCCAGCTAGACCTTTTACCCCAGGTCACCACACTTACCATTGAACAT
TGTCTCAATCTGGAATCCCTATGTATAGGCGAGAGACCTGTCCCTGCTCTCTGTCGTCTGACAATCAGTCACTGTCCTAATTTGGTGTCTTTCCCCAAAG
GAGGACTAGCTGCCCCGGATTTGACAAGCCTTGTGTTAGAGGGTTGCTTATATTTGAAGTCGTTGCCGGAAAACATGCGTTCCCTCCTACCTTCCCTACA
GGATTTGCAATTGATCTCACTTCCAGAAGTTGATTCGTTTCCTGAAGGAGGTTTACCCTCCAATTTAAACACACTTTGCATTGAAGATTGTATCAAACTC
AAGGTATGTGGTTTGCAAGCACTCTCTTCTCTTTCATACTTCGAATTTAGTGGGAACGATGTGGAGTCTTTTGACGAGGAGACGTTGCCCTCTACTCTCA
CAACCCTTAAAATCAAACGTCTGGGAAACCTGAAGTCTCTGGACTATAAGGGGCTTCATCACCTCACCTCTCTTCAGGGATTGGGTATCGAGGGTTGCCC
TAAGCTTGAGTCCATATCAGAGCAGGCGCTTCCCTCCTCCCTTGAATACCTTTATCTCAGAAATCTAGAATCTCTGGACTACGTGGGGCTTCATCACCTC
ACCTCTCTTGACACATTGAAGATCAAGAGTTGCCCTAAACTTGAGTTCGTATCAGAACAGGTGCTGCCCTCCTCCCTTGAATACCAGGGGCTTCATCACC
TCACCTCTCTTAGAAATTTGAGTATCGAGAGTTACCCTAAACTTGAGCACATATCAGAACAGGGGCTGCCCTCCTCCCTTGAATGCCTTCATCTCTGTAA
ACTGGAATCTTTGGACTACATTGGGCTTCAGCACCTCACCTCTCTTCACAAAATGAAGATCGGGAGTTGTCCTAAACTTGAATCGTTGCAGGGGCTGCCT
TCCTCCCTTGAATTCCTTCAGTTATGGGATCAGCAAGACCGGGACTATAAGGAACTTCGGCACCTCACCTCTCTTCGCAAAATGAATATAAGAAGGAGCC
TTAAGCTTGAGTACCTTCAAGAAGGCACACTTCCCTCCTCCCTTAAAGACCTTGAAATCCAGGATTTGGAAGATCTGGATTATAAGGGGTTTCGGCACCT
CAGCTCTCTTCGTAAATTACATATCTGTAATTCTCCTAAGCTCGAGTTCGTGCCAGGAGAGGAGCTGCCCTCCTCCCTTGTATCTCTTAAAATATCAGGT
CTGATAAATCTGAAATCTGTAATGAGGCTTCAGCACCTCACCTCTCTTCGCAAATTGATTATCCGGGATTGTCCTAAGCTTGAGTACTTGCCAACAGAAG
AGCTGTCCTTACCCCTAGTTCCTGATATTAGTGGTTGTCCTTTTGTAGAACCGAGTTGTTGCATTAGTTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.017G015200.1 pacid=42813172 polypeptide=Potri.017G015200.1.p locus=Potri.017G015200 ID=Potri.017G015200.1.v4.1 annot-version=v4.1
MALELIGGSILSALVEVLVDRLASREVLDSFKSHKLDDGLLEKLNETLNTVNGLLDDAEEKQITKRAVKNWLNDVKHAVYEAEDILDEINYEYLRSKDID
TPRPDSNWARNLVPFLNPANRRMKEIGAELQKIYEKLERLLKHKGDLRHIEGTGGWRPLSEKTTPLVNEPDVYGRDADKEAIMEHLLTQHKTDGPNLVVV
PIVGMGGIGKTTLAQLIYNDEKVDQFFQLKAWVWASQQFDVTRIIEDIIKKIKARTCPTKEPDESKEPNESLMEAVKGKKLLLVLDDAWNIEYNEWDKLL
LPLRYVEHGSKIVVTTREEDVAKVTQTVIPSHRLNVISDEDCWKLFARDAFSGVNSGAVSHLEEFGRVIVRKCKGLPLAAKTLGGLLHSVGDVKQWEKIS
NSSMWGSSNENIPPALTLSYYYLPSHLKRCFAYCAIFPKDYVFKKDRLITEWMAHGFLVQPRGVEEMEDIGEKYFNDLVSRSLFQQSTGDSFFSMHDLIS
DLAEYVSGEFCFKLGINESGSGLESEHSCSLPERTRYLSITSAAAYGGGLRIFRSIHGVQHLRALFPLKFFVEVDIEALNDILPNLKRLRMLSLCHPKDI
SSQLLNSIGNLKHLRHLDLSQTVFKRLPESVCTLYYLQSLLLKECRLLMELPSNLSNLVDLQHLDIEGTNLKEMPPKMGKLTKLRILESYIVGKDSGSSM
KELGKLSHIRKKLSIRNLRDVANAQDALDANLKGKKKIEELGLTWDGSTDDTPHERDVLEKLEPSEDVKELAIIGYGGTTFPGWLGNSSFSNMVTLLLSG
CTNCILLPPLGQLPSLEELEIEGFDEVVAVGSEFYGSDPPMEKPFKSLITLKFEGMKKWQEWNTDVAGAFPHLENLLIAGCPELTNGLPNHLPSLLILEI
RACPQLVVSIPEAPLLREINVSEGDESRITGRTSSYYCYYLPDQPPKCLHFRRDPQLTGMEQMSHLDPRLFTDIKIEECSSFKCCQLDLLPQVTTLTIEH
CLNLESLCIGERPVPALCRLTISHCPNLVSFPKGGLAAPDLTSLVLEGCLYLKSLPENMRSLLPSLQDLQLISLPEVDSFPEGGLPSNLNTLCIEDCIKL
KVCGLQALSSLSYFEFSGNDVESFDEETLPSTLTTLKIKRLGNLKSLDYKGLHHLTSLQGLGIEGCPKLESISEQALPSSLEYLYLRNLESLDYVGLHHL
TSLDTLKIKSCPKLEFVSEQVLPSSLEYQGLHHLTSLRNLSIESYPKLEHISEQGLPSSLECLHLCKLESLDYIGLQHLTSLHKMKIGSCPKLESLQGLP
SSLEFLQLWDQQDRDYKELRHLTSLRKMNIRRSLKLEYLQEGTLPSSLKDLEIQDLEDLDYKGFRHLSSLRKLHICNSPKLEFVPGEELPSSLVSLKISG
LINLKSVMRLQHLTSLRKLIIRDCPKLEYLPTEELSLPLVPDISGCPFVEPSCCIS
|
DESeq2's median of ratios [POPLAR]
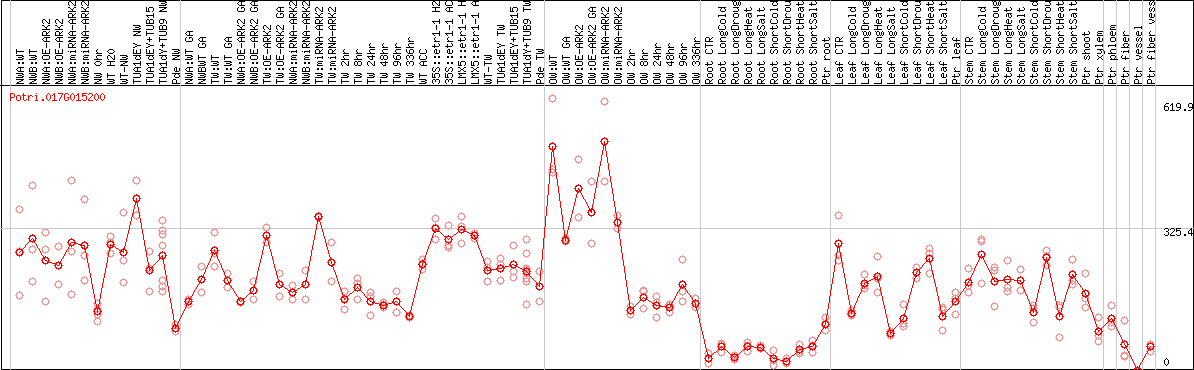
Coexpressed genes
Potri.017G015200 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
