Potri.017G015600 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.017G015600.5 pacid=42813858 polypeptide=Potri.017G015600.5.p locus=Potri.017G015600 ID=Potri.017G015600.5.v4.1 annot-version=v4.1
ATGGCTGTGGAGTACATTGGACGATCAATTCTCTTTGCTGTCATTGAGGTTCTGGGCGAGAAGTTGACCACTCCTGAGATTCTGGGCTTCTTCAAAAGCC
ACAAGCTCAATGATGGTCTTCTCGGAAAGTTGAAAGAAACACTGAATACTCTCAACGGACTTCTCGACGACGCGGAGGAGAAGCAGATTACCAAGCCTGC
TGTGCAGAGGTGGCTTAATGATGCTAGACATGCTGTGTATGAAGCTGAGGACTTAATGGAGGAGATTGAATATGAACATCTACGATCCAAGGATATTAAA
GCTGCCTCTCGACGAGTTAGGAATCGGGTAAGGAATTTGTTTCCAATTCTTAATCCAGCTAATAAAAGGATGAAAGAGATGGAAGCAGGGTTACAAAAGA
TCTATGAGAAGCTTGAACGTTTAGTTAAACACAAGGGTGATCTGCGCCACATAGAAGGTAATGGTGGAGGGAGACCATTGTCAGAGAAAACAACTCCTGT
GGTTGATGAATCTCATGTATATGGAAGGGAAGCTGATAAGGAAGCCATAATGAAGTACTTGTTGACAAAAAACAATACAAATGGCGCAAATGTAGGTGTG
ATTCCTATTGTGGGCATGGGAGGGGTTGGTAAAACCACTCTTGCTCAGCTTATCTACAAAGACAGAAGGGTAGATAAGTGCTTCGAGCTCAAAGCCTGGG
TCTGGGCTTCTCAACAATTTGACGTCACCAGGATAGTTGATGATATTCTTAAGAAGATCAACGCAGGTACTTGCGGCACCAAAGAACCAGATGAATCTCT
AATGGAGGCAGTGAAAGGGAAAAAACTTCTACTTGTTCTAGATGATGCGTGGAACATTGTGTACAATGAATGGGTTAAATTACTGCTGCCTTTGCAGTAT
GCGGAGCCTGGAAGTAAGATTGTCGTGACAACACGGAATGAAGATGTAGCAAAAGTCACGCAAACTGTCATCCCTTCTCACCATTTGAAGGGAATTAGCG
ATGAAGATTGCTGGCAGTTGTTTGCAAGGCATGCATTTAGCGGTGCAAATTCTGGAGCAGTCTCGCACTTGGAAACATTTGGCAGAGAAATAGCGAGAAA
GTGCAAAGGTTTACCGCTGGCAGCGAAAACTCTTGGGGGTCTTTTGCACTCTGTAGGAGATGTCAAGCAATGGGAGAAGATATCCAAGAGCAGGATGTGG
GGTTTATCGAATGAAAACATCCCTCCAGCTCTAACATTGAGCTATTATTATCTCCCTTCACACCTGAAACGTTGTTTTGCTTATTGTGCAATATTTCCCA
AGGGTTATGTATTTGAGAAGAATCAAGTAATCACTTCATGGATGGCACAAGGTTTTTTAGTCCAGTCCAGAGGCGTTGAGGAGATGGAAGAAATAGGTGA
CAAATACTTCAATGACCTTGTCTCCAGGTCATTGTTTCAGCAATCACTTTACGCCCCATCATATTTCAGCATGCACGACCTCACAAGTGATCTAGCTGAA
TACATGTCAGGAGAATTCTGCTTTAAATTTGTTATGGATGGAGAATCGGGCTCTGGGTTGGAGGGTGAAAATTCATGCACGCTTCCAGAAAGCACTCGTC
ATTTATCAATCACTTCAACACTATATGATGGGGTCTCTAAGATATTTCCTCGCATTCATGGAGTCCAACATTTGCGCACCTTGTCTCCACTAACCTACGT
GGGGGGGATTGATAGTGAGGTGCTGAATGACATGTTGACAAATCTTAAGCGCTTGCGAACGCTATCTTTATATCGGTGGAGCTATAAATCTTCTCGGTTG
CCCAATTCCATCGGCAACTTGAAACATTTGCGGCACTTGGACCTTTCTCAGACATTAATTAAACGGTTGCCCGAAAGTGTGAGCACCTTGTACTATTTGC
AAACTCTATTGTTAAGAGAGTGTCGACATCTCATGGAGTTGCCATCCAACATTTCGAACTTGGTCGACTTACAACATCTTGATATTGAAGGGACAAATTT
GAAAGAGATGCCACCAAAAATGGGGAAACTCACAAAGCTTCGAACTTTGCAATATTACATTGTGGGAAAAGAGAGTGGGTCTAGCATGAAAGAGCTGGGG
AAGCTCTCACATATAAGGAAAAAACTTTCTATTCGGAATCTCAGAGATGTTGCAAATGCTCAAGATGCTTTGGATGCTAATTTGAAGGGTAAGAAGAAGA
TTGAGAAGCTGAGGTTGATATGGGTTGGCAATACGGATGACACACAACATGAGAGAGACGTGCTTGAGAAATTGGAGCCTTCTGAAAATGTGAAACAGCT
TGTCATTACTGGTTATGGGGGTACGATGTTTCCAGGTTGGTTTGGAAACTCTTCCTTCTCAAATATGGTAGCGTTGACACTTTCTGGATGTAAGAACTGC
ATCTCCTTACCACCACTGGGGCAGCTTTCATCTCTAGAAGAGCTCCAAATTAAAGGATTTGATGAAGTCGTGGCAGTTGACTCTGAGTTCTATGGAAGTG
ACTCTTCGATGGAGAAGCCATTTAAATCCCTCAAAATATTAAAGTTCGAGGGGATGAAAAAATGGCAGGAATGGAATACAGATGTAGCTGCTGCTTTCCC
TCATCTTGCAAAGCTCTTGATAGCAGGTTGTCCCGAGTTAACAAATGGCTTGCCTAATCACCTTCCTTCTTTATTGATACTTGAGATTAGAGCGTGTCCA
CAGCTTGTGGTTTCAATTCCAGAGGCTCCCCTGCTCACCGAAATCAATGTATTTGATGGTTCTAGTGGTCGTATAAATGCATCGGTATTATATGGCGGTG
GGCGGTGCCTTCAATTTAGAGAATATCCCCAACTGAAGGGAATGGAGCAAATGAGTCATGTGGATCCCTCTTCTTTCACAGATGTTGAAATCGATAGATG
TAGTTCATTTAACTCCTGCCGGCTAGACCTTTTACCCCAGGTATCCACACTTACAGTCAAACAATGTCTCAATCTCGAATCTCTTTGTATAGGCGAGAGA
TCTCTTCCTGCTCTCCGTCATTTGACTGTCAGACACTGTCCTAATTTGGTGTCTTTCCCCGAAGGAGGACTAGCTGCCCCAGATTTGACAAGCCTTGTGT
TAGAGGGTTGCTTATATTTGAAGTCTTTGCCAGAAAACATGCATTCCCTCCTCCCTTCCCTTGAGGATTTGCAATTGAGATCACTTCCAGAAGTTGATTC
GTTTCCCGAAGGAGGTTTACCCTCCAAATTACACACACTTTGCATTGTGGATTGTATCAAGCTCAAGGTATGTGGTTTGCAAGCACTCCCTTCTCTTTCA
TGCTTCAGATTTACTGGGAACGATGTGGAGTCTTTTGACGAGGAGACGTTGCCCTCTACTCTCAAAACCCTTAAAATCAAACGTCTGGGAAATCTGAAGT
CTCTGGACTATAAGGGGCTTCATCACCTCACCTCTCTTAGGAAATTGAGTATCGAGGGTTGCCCTAAGCTTGAGTCCATATCAGAGCAGGCGCTTCCCTC
CTCCCTTGAATGCCTTCATCTCATGACACTGGAATCTCTGGACTACATGGGGCTTCAGCACATCACCTCTCTTCGCAAATTGAAGATCTGGAGTTGTCCA
AAGCTTGCATCGTTGCAGGGGCTGCCTTCTTCCCTTGAATGCCTTCAATTATGGGATCAGCGAGGTCGGGACTCCAAGGAGCTTCAGCACCTCACCTCTC
TTCGCACATTGATTCTCAAGAGTCCTAAGCTGGAGTCCCTTCCAGAAGAAATGCTTCCCTCCTCCCTTGAAAACCTTGAAATCTTGAATCTTGAAGATCT
GGAATATAAGGGGCTTCGGCACCTCACCTCCCTTCGTAAATTACGTATCTCTAGTTCTCCCAAGCTTGAGTCCGTGCCAGGTGAGGGGCTGCCCTCCTCC
CTTGTATCTCTTCAAATCTCAGATCTCAGAAATCTTAAATCTCTGAACTATATGGGGCTTCAACACTTCACCTCTCTTCGCAAATTGTTGATCTCGCATT
CTCCTAAGCTTGAGTTCATGCCAGAAGAAGGGCTGCCCCCCTCCCTTGAATATCTCAAGATTATCGATTGTCCTTTGCTGGCCACAAGGTGTGAAAGGGA
GACAGGGGAAGATTGGCCCAAAATTTCTCACATCCGCAGTATAAAAATTTCTAATTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.017G015600.5 pacid=42813858 polypeptide=Potri.017G015600.5.p locus=Potri.017G015600 ID=Potri.017G015600.5.v4.1 annot-version=v4.1
MAVEYIGRSILFAVIEVLGEKLTTPEILGFFKSHKLNDGLLGKLKETLNTLNGLLDDAEEKQITKPAVQRWLNDARHAVYEAEDLMEEIEYEHLRSKDIK
AASRRVRNRVRNLFPILNPANKRMKEMEAGLQKIYEKLERLVKHKGDLRHIEGNGGGRPLSEKTTPVVDESHVYGREADKEAIMKYLLTKNNTNGANVGV
IPIVGMGGVGKTTLAQLIYKDRRVDKCFELKAWVWASQQFDVTRIVDDILKKINAGTCGTKEPDESLMEAVKGKKLLLVLDDAWNIVYNEWVKLLLPLQY
AEPGSKIVVTTRNEDVAKVTQTVIPSHHLKGISDEDCWQLFARHAFSGANSGAVSHLETFGREIARKCKGLPLAAKTLGGLLHSVGDVKQWEKISKSRMW
GLSNENIPPALTLSYYYLPSHLKRCFAYCAIFPKGYVFEKNQVITSWMAQGFLVQSRGVEEMEEIGDKYFNDLVSRSLFQQSLYAPSYFSMHDLTSDLAE
YMSGEFCFKFVMDGESGSGLEGENSCTLPESTRHLSITSTLYDGVSKIFPRIHGVQHLRTLSPLTYVGGIDSEVLNDMLTNLKRLRTLSLYRWSYKSSRL
PNSIGNLKHLRHLDLSQTLIKRLPESVSTLYYLQTLLLRECRHLMELPSNISNLVDLQHLDIEGTNLKEMPPKMGKLTKLRTLQYYIVGKESGSSMKELG
KLSHIRKKLSIRNLRDVANAQDALDANLKGKKKIEKLRLIWVGNTDDTQHERDVLEKLEPSENVKQLVITGYGGTMFPGWFGNSSFSNMVALTLSGCKNC
ISLPPLGQLSSLEELQIKGFDEVVAVDSEFYGSDSSMEKPFKSLKILKFEGMKKWQEWNTDVAAAFPHLAKLLIAGCPELTNGLPNHLPSLLILEIRACP
QLVVSIPEAPLLTEINVFDGSSGRINASVLYGGGRCLQFREYPQLKGMEQMSHVDPSSFTDVEIDRCSSFNSCRLDLLPQVSTLTVKQCLNLESLCIGER
SLPALRHLTVRHCPNLVSFPEGGLAAPDLTSLVLEGCLYLKSLPENMHSLLPSLEDLQLRSLPEVDSFPEGGLPSKLHTLCIVDCIKLKVCGLQALPSLS
CFRFTGNDVESFDEETLPSTLKTLKIKRLGNLKSLDYKGLHHLTSLRKLSIEGCPKLESISEQALPSSLECLHLMTLESLDYMGLQHITSLRKLKIWSCP
KLASLQGLPSSLECLQLWDQRGRDSKELQHLTSLRTLILKSPKLESLPEEMLPSSLENLEILNLEDLEYKGLRHLTSLRKLRISSSPKLESVPGEGLPSS
LVSLQISDLRNLKSLNYMGLQHFTSLRKLLISHSPKLEFMPEEGLPPSLEYLKIIDCPLLATRCERETGEDWPKISHIRSIKISN
|
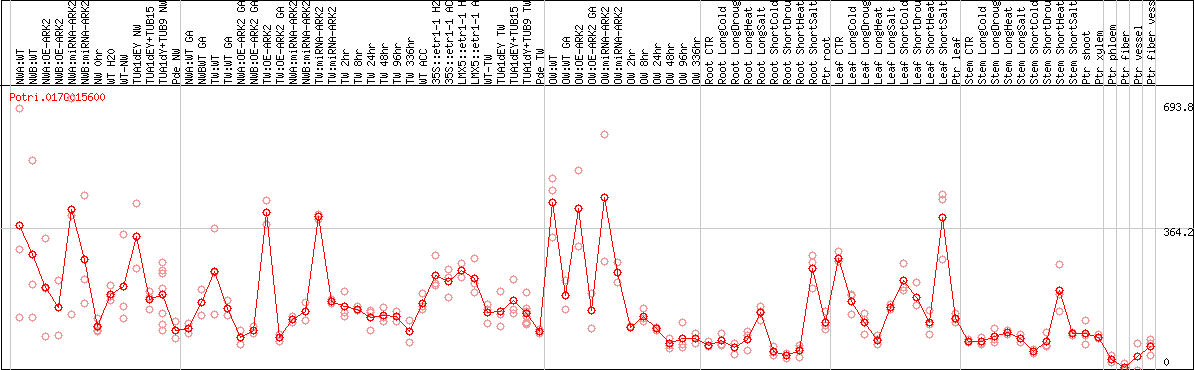
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.017G015600 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
