Potri.017G105201 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.017G105201.1 pacid=42814257 polypeptide=Potri.017G105201.1.p locus=Potri.017G105201 ID=Potri.017G105201.1.v4.1 annot-version=v4.1
ATGGCTTCCAACAGCGTGCAAGGAATAACCTCATCTTCATCTCCTCCTCCTCCGCTATATATGTATGATGTGTTCCTTAGTTTTAGAGGCAAAGACACCC
GGAACAACTTTACTAGTCATCTATATTCTAATCTGAAACAGAGAGGCATCGATGTATACATGGATGATAGGGAGCTCGAGAGAGGAAAGACCATCGAAAC
TGCTCTCTGGAAGGCCATCGAAGAATCAAGGTTTTCTGTCATTATATTTTCAAGAGATTACGCTTCTTCACCATGGTGCTTGGATGAACTTGTCAAGATT
GTTCAGTGCATGAAAGAGATGGGGCAAACTGTTCTGCCAGTTTTTTATGATGTGGATCCCTCAGAGATGACCGAGCGGAAACGGAAATATGAGGAAGCCT
TCGTTGAGCATGAACAAAATTTCAAGGAAAACTTGGAGAAGGTCGGGAACTGGAAAGATTGTCTCTCTACGGTGGCCAATCTATCTGGCTGGGATGTCCG
GAATAGGAATGAATCGAAATCAATTAAAATAATTGCTGAATACATATCGTACAAACTAAGCGTCACTCTGCCAACAATTAGCAAAAAATTAGTTGGAATA
GATTCTCGTGTAGAGGTATTAAATGGCTATATAGGTGAAGAAGTAGGCAAAGCAATCTTCATAGGGATTTGTGGAATGGGTGGCATAGGTAAGACGACTG
TTGCAAGGGTACTATATGATAGGATTCGTTGGCAATTTGAAGGCAGCTGCTTCTTAGCAAATGTCAGGGAAGTTTTTGCTGAGAAAGATGGACCACGCCG
TTTACAGGAACAACTTCTTTATGAAATCTTAATAGAACGTGCTTTAGTATGGGATTCTTCTAGAGGGATTGAGATGATAAAGCGGAGGTTACGACTTAAA
AAGATTCTTCTTATCCTTGATGATGTAGATGACAAAAAACAACTAGAATTTTTGGCTGCGGAGCGTGGATGGTTTGGTCCAGGGAGCAGAATTATCATAA
CAAGCAGAGATACAAATGTGTTTACTGGAAATGATGATACTAAAATTTATGAGGCTGAGAAATTGAATGATGATGATGCTCTTATGTTGTTTAGCCAGAA
AGCTTTCAAAAATGACCAACCTGCTGAAGATTTTGTGGAACTATCCAAGCAAGTTGTGGGCTATGCTAATGGCCTTCCACTGGCTCTTGAAGTTATAGGT
TCGTTTTTGTATGGAAGAAGTATCCCTGAATGGAGAGGTGCAATTAATAGAATGAATGAGATTCCTGACTGCAAAATCATTGATGTGCTTCGCATTAGTT
TTGATGGTCTCCATGAATCAGATAAGAAAATATTTTTAGACATTGCCTGTTTCCTGAAGGGGTTTAAAATTGATCGAATAACAAGGATACTTGACAGCCG
TGGATTCCATGCAGGTATTGGAATACCAGTTCTTATTGAGAGATCTCTCATAAGTGTCTCTCGGGATCAAGTCTGGATGCATAATTTATTACAGATAATG
GGTAAGGAAATTGTTCGTTGTGAATCCCCTGAAGAGCCTGGAAGGCGCAGTAGATTGTGGACATACGAGGATGTCTGCCTTGCACTGATGGACAACACAG
GAAAAGAAAAAATAGAAGCCATATTCTTGGACATGCCTGGAATAAAAGAGTCACAATGGAACATCGAAGCCTTCTCTAAAATGAGCAGACTAAGATTGCT
CAAAATCAACAACGTGCAGCTTTCTGAAGGACCTGAAGATCTTTCCAATAAGTTACAATTTCTTGAATGGCATTCTTACCCTTCAAAATCATTGCCAGTT
GGTTTGCAGGTGGATCAGCTAGTTGAACTTCACATGGCTAACAGTAATCTTGAGCAATTATGGTATGGGTGTAAGAGTGCAGTTAATTTGAAAATCATCA
ATCTCAGCAACTCACTATACCTGACCAAGACTCCAGATCTTACTGGAATTCCAAATCTCGAGAGTCTGATTCTTGAAGGTTGTACAAGTTTGTCTGAGGT
CCATCCATCACTTGCACATCACAAGAAGCTTCAATATATGAACCTTGTGAACTGCAAAAGCATTAGGATTCTCCCGAACAATCTAGAAATGGGATCACTT
AAAGTTTGCATTCTTGATGGCTGCTCAAAACTTGAGAAGTTTCCAGATATAGTAGGAAACATGAAATGTTTGATGGTGCTTCGATTGGATGGGACTGGTA
TTACAAAACTATCTTCATCGATGCATCATTTGATTGGCTTAGGTCTATTGAGCATGAACAGCTGCAAGAACCTTGAAAGCATTCCAAGTAGCATAGGTTG
TTTGAAATCCCTTAAAAAACTTGATCTATCTGGCTGCTCTGAACTTAAATATATACCAGAGAAGTTGGGGGAAGTTGAAAGTTTGGAGGAGTTTGATGTA
AGTGGAACTTCAATAAGACAACTGCCAGCATCCATTTTTCTTTTAAAGAATCTTAAAGTATTATCTTTGGATGGATTCAAAAGAATAGTTATGCCGCCTT
CTTTGTCTGGTCTGTGTTCTTTAGAAGTACTAGGTTTATGCGCTTGCAATCTAAGAGAAGGGGCACTTCCTGAAGATATTGGCTGCTTATCTTCATTGAG
GTCCTTAGATCTGAGCCAGAATAACTTTGTTAGCCTGCCTAAAAGCATAAATCAGCTTTTTGAACTGGAAATGCTTGTCTTGGAGGATTGCACGATGCTT
GAATCATTGCCTAAGGTTCCATCTAAAGTTCAAACGGTATGTTTGAATGGTTGTATAAGCCTAAAAACAATTCCAGATCCGATAAACCTAAGCAGTTCCA
AAATCTCAGAGTTCGTATGTCTTAACTGCTGGGAATTGTACAATCATTATGGCCAAGACAGCATGGGGTTGACCCTGCTTGAAAGATACTTCCAGGGTTT
GTCTAATCCAAGACCTGGATTTGGCATCGCCATTCCAGGAAATGAAATTCCAGGCTGGTTTAACCATCAAAGTAAGGGATCTTCAATTAGTGTGCAAGTG
CCTAGTTGGAGCATGGGGTTTGTTGCGTGTGTTGCATTCGGTGTAAATGGTGAAAGTCCTTCTCTTTTTTGTCATTTCAAAGCCAATGGAAGAGAGAATT
ATCCTTCATCGCCAATGTGTATTAGTTGTAACTCTATCCAAGTTCTGTCAGATCATATTTGGCTATTCTATCTATCTTTTGATTACCTTAAAGAGCTTCA
AGAGTGGCAGCATGGATCCTTTAGCAACATTGAGTTGTCGTTTCATTCTTCCCAGCCAGGAGTGAAGGTGAAGAATTGTGGTGTCCGTTTGCTATCTTCT
ATATATATTACACCACAACTATCTTCTGCCCATTTTATTGTTACAAGCAAAGAAGTAGCTTCTTCATTTAAAGCTTCTTTAGCTTTTTCTTCGTCGTACC
ATCAATGGAAGGCTAATGTTTTCCCTGGCATCAGAGTCGCAGACACTAATCTAGCCCTGAGATTCATTGTGCCAGTCGAGAAGGAACCAGAGAAAGTAAT
GGCAATTAGATCAAGACTCTTTGAGGCCATTGAAGAATCAGGGCTGTCAATTATTATATTTGCAAGAGATTGTGCTTCTTTACCATGGTGCTTTGAAGAA
CTTGTCAAGATTGTCGGATTCATGGATGAGATGAGATCGGACATTGTGTTTCCAGTTTCTCGTGATGTCAAACAATCAAAGATAGATGATCAAACAGAGA
GTTACACAATTGTCTTTGATAAGAATGAAGAAAATCTCAGGGAAAACGAGGAGAAGGGACAAAGATGGATGGATATTCTCACTAAAGTTGAGATTTCATC
TGGATCGAATTCATTGAAAAGGTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.017G105201.1 pacid=42814257 polypeptide=Potri.017G105201.1.p locus=Potri.017G105201 ID=Potri.017G105201.1.v4.1 annot-version=v4.1
MASNSVQGITSSSSPPPPLYMYDVFLSFRGKDTRNNFTSHLYSNLKQRGIDVYMDDRELERGKTIETALWKAIEESRFSVIIFSRDYASSPWCLDELVKI
VQCMKEMGQTVLPVFYDVDPSEMTERKRKYEEAFVEHEQNFKENLEKVGNWKDCLSTVANLSGWDVRNRNESKSIKIIAEYISYKLSVTLPTISKKLVGI
DSRVEVLNGYIGEEVGKAIFIGICGMGGIGKTTVARVLYDRIRWQFEGSCFLANVREVFAEKDGPRRLQEQLLYEILIERALVWDSSRGIEMIKRRLRLK
KILLILDDVDDKKQLEFLAAERGWFGPGSRIIITSRDTNVFTGNDDTKIYEAEKLNDDDALMLFSQKAFKNDQPAEDFVELSKQVVGYANGLPLALEVIG
SFLYGRSIPEWRGAINRMNEIPDCKIIDVLRISFDGLHESDKKIFLDIACFLKGFKIDRITRILDSRGFHAGIGIPVLIERSLISVSRDQVWMHNLLQIM
GKEIVRCESPEEPGRRSRLWTYEDVCLALMDNTGKEKIEAIFLDMPGIKESQWNIEAFSKMSRLRLLKINNVQLSEGPEDLSNKLQFLEWHSYPSKSLPV
GLQVDQLVELHMANSNLEQLWYGCKSAVNLKIINLSNSLYLTKTPDLTGIPNLESLILEGCTSLSEVHPSLAHHKKLQYMNLVNCKSIRILPNNLEMGSL
KVCILDGCSKLEKFPDIVGNMKCLMVLRLDGTGITKLSSSMHHLIGLGLLSMNSCKNLESIPSSIGCLKSLKKLDLSGCSELKYIPEKLGEVESLEEFDV
SGTSIRQLPASIFLLKNLKVLSLDGFKRIVMPPSLSGLCSLEVLGLCACNLREGALPEDIGCLSSLRSLDLSQNNFVSLPKSINQLFELEMLVLEDCTML
ESLPKVPSKVQTVCLNGCISLKTIPDPINLSSSKISEFVCLNCWELYNHYGQDSMGLTLLERYFQGLSNPRPGFGIAIPGNEIPGWFNHQSKGSSISVQV
PSWSMGFVACVAFGVNGESPSLFCHFKANGRENYPSSPMCISCNSIQVLSDHIWLFYLSFDYLKELQEWQHGSFSNIELSFHSSQPGVKVKNCGVRLLSS
IYITPQLSSAHFIVTSKEVASSFKASLAFSSSYHQWKANVFPGIRVADTNLALRFIVPVEKEPEKVMAIRSRLFEAIEESGLSIIIFARDCASLPWCFEE
LVKIVGFMDEMRSDIVFPVSRDVKQSKIDDQTESYTIVFDKNEENLRENEEKGQRWMDILTKVEISSGSNSLKR
|
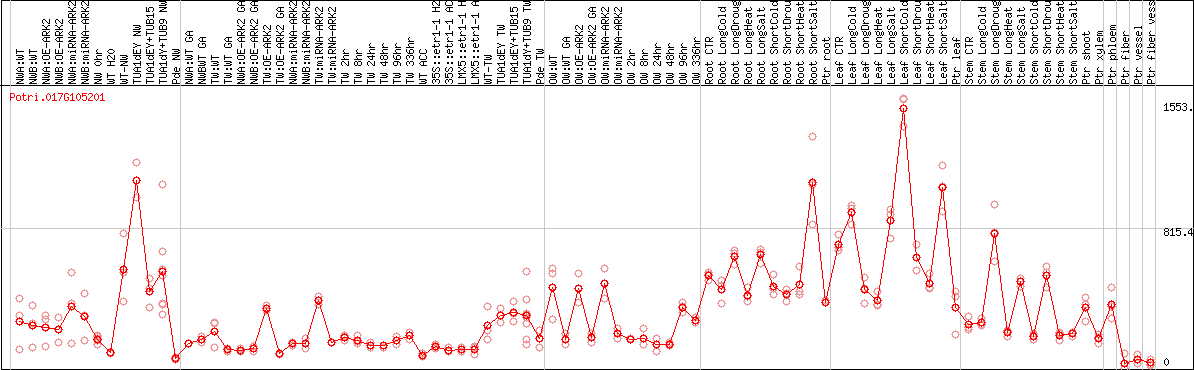
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.017G105201 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
