External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G25060 79 / 1e-18
AtENODL14
early nodulin-like protein 14 (.1)
AT3G20570 77 / 4e-18
AtENODL9
early nodulin-like protein 9 (.1)
AT5G20230 73 / 2e-16
SAG14, ATBCB
SENESCENCE ASSOCIATED GENE 14, BLUE COPPER BINDING PROTEIN, blue-copper-binding protein (.1)
AT4G31840 70 / 2e-15
AtENODL15
early nodulin-like protein 15 (.1)
AT2G32300 71 / 4e-15
UCC1
uclacyanin 1 (.1)
AT3G18590 66 / 8e-14
AtENODL5
early nodulin-like protein 5 (.1)
AT5G53870 65 / 7e-13
AtENODL1
early nodulin-like protein 1 (.1)
AT1G48940 63 / 9e-13
AtENODL6
early nodulin-like protein 6 (.1)
AT2G31050 60 / 2e-11
Cupredoxin superfamily protein (.1)
AT4G28365 60 / 2e-11
AtENODL3
early nodulin-like protein 3 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.018G129000
255 / 8e-88
AT3G20570 76 / 7e-17
early nodulin-like protein 9 (.1)
Potri.018G128800
213 / 7e-72
AT2G25060 72 / 1e-16
early nodulin-like protein 14 (.1)
Potri.018G129200
92 / 2e-23
AT2G32300 94 / 9e-23
uclacyanin 1 (.1)
Potri.018G129400
90 / 6e-23
AT5G20230 114 / 6e-32
SENESCENCE ASSOCIATED GENE 14, BLUE COPPER BINDING PROTEIN, blue-copper-binding protein (.1)
Potri.006G067400
87 / 6e-22
AT5G20230 112 / 2e-31
SENESCENCE ASSOCIATED GENE 14, BLUE COPPER BINDING PROTEIN, blue-copper-binding protein (.1)
Potri.006G067200
85 / 9e-22
AT5G20230 61 / 3e-12
SENESCENCE ASSOCIATED GENE 14, BLUE COPPER BINDING PROTEIN, blue-copper-binding protein (.1)
Potri.001G192100
82 / 9e-20
AT5G20230 100 / 2e-26
SENESCENCE ASSOCIATED GENE 14, BLUE COPPER BINDING PROTEIN, blue-copper-binding protein (.1)
Potri.006G067300
81 / 9e-19
AT5G20230 97 / 2e-24
SENESCENCE ASSOCIATED GENE 14, BLUE COPPER BINDING PROTEIN, blue-copper-binding protein (.1)
Potri.011G135400
72 / 8e-16
AT3G20570 138 / 4e-41
early nodulin-like protein 9 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10006657
96 / 6e-24
AT1G45063 106 / 1e-26
copper ion binding;electron carriers (.1.2)
Lus10035911
87 / 2e-21
AT5G20230 95 / 3e-24
SENESCENCE ASSOCIATED GENE 14, BLUE COPPER BINDING PROTEIN, blue-copper-binding protein (.1)
Lus10038098
88 / 5e-21
AT1G45063 107 / 1e-27
copper ion binding;electron carriers (.1.2)
Lus10025752
82 / 1e-19
AT5G20230 101 / 1e-26
SENESCENCE ASSOCIATED GENE 14, BLUE COPPER BINDING PROTEIN, blue-copper-binding protein (.1)
Lus10008720
79 / 8e-19
AT1G72230 140 / 2e-42
Cupredoxin superfamily protein (.1)
Lus10011158
76 / 2e-17
AT3G20570 149 / 2e-45
early nodulin-like protein 9 (.1)
Lus10020944
74 / 8e-17
AT1G72230 141 / 1e-42
Cupredoxin superfamily protein (.1)
Lus10043063
74 / 9e-17
AT3G20570 153 / 4e-47
early nodulin-like protein 9 (.1)
Lus10019405
74 / 5e-16
AT1G45063 107 / 3e-27
copper ion binding;electron carriers (.1.2)
Lus10002614
70 / 8e-16
AT2G32300 100 / 1e-26
uclacyanin 1 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0026
CU_oxidase
PF02298
Cu_bind_like
Plastocyanin-like domain
Representative CDS sequence
>Potri.018G128900.1 pacid=42801184 polypeptide=Potri.018G128900.1.p locus=Potri.018G128900 ID=Potri.018G128900.1.v4.1 annot-version=v4.1
ATGGCTAGCCGATTGTGTTTCAATATTGGGTTCTTGATTGTTGCATCGGCGGGTTTGCTTCACGGCGCATATGCAGCTAATACTTACACGGTTGGAGGTG
ATTTGGGATGGATTGTTCCTCCGAACAGCTCTTACTATGAGGAATGGACCAGCCAGAGCACATTTCAGATAGGAGACTCCTTCGTATTCAACTGGACAAC
CGGAACACATACCGCAACTGAAGTAAGTACCAAGGAGGAGTATGATAATTGCACAAAGATGGGAATAATATTGAAAGATGCAGGAGTTAAAGTCACTTTC
AATGCAAATGGTACGCACTACTTCCTCTGCTCTGAAGGAACACACTGTGAGCAAGGCCAGAAGATGATAATCAAGATTGGAGATGGCATACCGCCAAGTT
TTGCAGCTCCATCTCTTACCGCGGCCGCCGCCTTATCTGCTCTTTTCTTCTCCACCTTAGCCATCTTCTTCTTAAACTAA
AA sequence
>Potri.018G128900.1 pacid=42801184 polypeptide=Potri.018G128900.1.p locus=Potri.018G128900 ID=Potri.018G128900.1.v4.1 annot-version=v4.1
MASRLCFNIGFLIVASAGLLHGAYAANTYTVGGDLGWIVPPNSSYYEEWTSQSTFQIGDSFVFNWTTGTHTATEVSTKEEYDNCTKMGIILKDAGVKVTF
NANGTHYFLCSEGTHCEQGQKMIIKIGDGIPPSFAAPSLTAAAALSALFFSTLAIFFLN
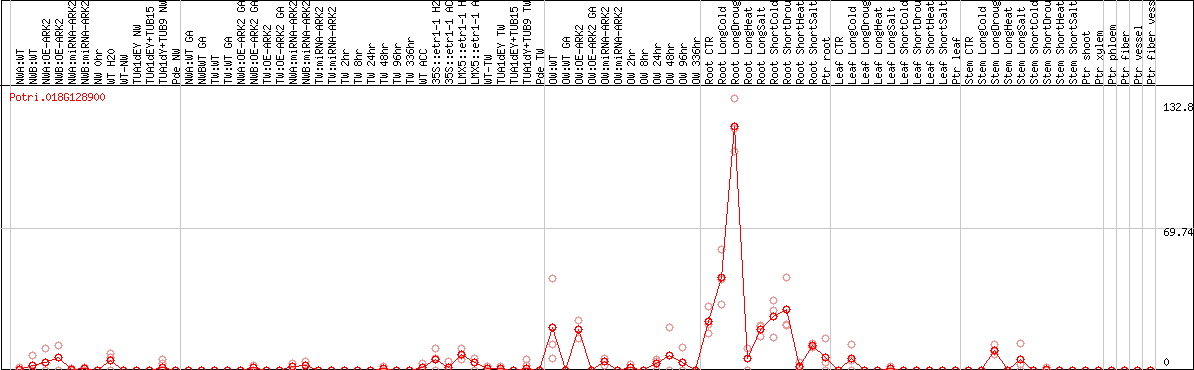
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.018G128900 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.