Potri.018G145532 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.018G145532.1 pacid=42800318 polypeptide=Potri.018G145532.1.p locus=Potri.018G145532 ID=Potri.018G145532.1.v4.1 annot-version=v4.1
ATGACGTGCAAGTTTTGTGGGCATTGTTTTACCCAAGGTACTTCCAGTAGGAGGATCAAATTGCATTTGGCTGGAGTGAGAGGGCGTGGTGTTAAAATTT
GTAAAAATGTTCCACAAGAAGTTCGAGACGCAGCTTGTGAAGCAGTTAATGATAGCCCTCCAGAAAAGAAACTTGAAACTGCTGCAGGCTCAAGCACAAA
CGAGGTCGCTAATGCAATTTCAGCTTCAACGCAAGAACAGAACATTGAAGTGACATATGTAGAGATGGCACTACAAGGAGGACCTTTTTTCACTAGAGAA
CTTGCATGGGCAAATGATTTGAGAGGTAGCTCTCATGATTCATCCCGACCAGCAGATGATCAACTCTGTTCATCAGTAAACAATGATGTCAATATGAAAG
ATGTGCAGAACAAGGTTGGAGTCAAGACAGAACCAGTACTGTTTCAAGTGGTGGAACAAAGCAATGCAGAACCTGTCAGTTTGGCAGGGCATGCTGGAAG
TATACAAGTAGGAGTTCATGGCATGGAACAAGGTGCGGAGGAAGAGAGAATTTGCCCGAACTTAGCGGCAAATGGCATGGAAAACCCTTGTGATGGATCT
AGTCAGCATTCCAGGAGACTCATAATTGATGCACAAGATAATACAAACTCTGCTTTTTTAATTAAGGAGGAGGATGTGGAGAACAATCTTGGAAGATTAG
TGCAGCCCGGCGCAGGAGCTAGCTCTTCTGGAGGGGTTGCATGCAAAACAAATAAGATTAAAGGAGATGCATTACTAACGAGAAAAATGGTGGGTCAAGC
ATTCAACGACCACAAGGAGACTATCCAATCTTTGTTGGAGCACAATGAAGTTTCAAGCATTGGCATCTATGGGATGGGCGGAGTGGGAAAGACCACATTG
GTGACACACATTTATAATCAGCTTCTAGAAAGACGAGACACTCATGTTTACTGGATTACTGGTTCACAGGATACCAGCATTAATAGATTGCAGACAAGTC
TCGCAAGACGTATTGGTTTAGATCTTTCCAGTGAAGATGAGGAGCTGCATAGAGCGGTCGCGTTGAAAAAAGAATTAATGAAGAAACAGAAATGGGTTCT
GATTTTAGATGATTTGTGGAAGGCTTTTGACCTACAAAAGCTAGGAGTTCCTGACCAAGTTGAGGGATGCAAGCTGATTCTCACAACTCGATCAGAAAAG
GTTTGTCAACAGATGAAAACCCAACACACGATCAAAGTGCAGCCTATTTCGGAGGAGGAGGCTTGGACCTTGTTTATTGAGAGACTTGGAGATGACATAG
CACTTTCTTCAGAAGTGAAACGAATTGCTGTGGATATTGTAAGGGAATGTGCTGGTTTGCCATTGGGAATTATTACAATGGCAAGAAGCATGAGGGGAGT
GGATGATCCATATGAGTGGACGGATACATTGAAGAAATTGAAAGAATCAAAATGTAGGGAAATGGAAGATGAGGTATTCCAGTTATTGAGGTTTAGTTAT
GATCAGTTAAATGATTTAGCATTACAACAATGTCTCTTGTATTGTGCATTGTATCCTGAAGATCATGAGATTGAAAGGGAAGAGCTGATAGGTTATTTGA
TCGATGAGGAAATAATTGAAGAAATGAGGAGCAGGCAAGCAGCATTTGACGAGGGACACATGATGCTTGATAAACTTGAAAAAGTCTGTTTATTGGAAAG
AGTTGGTTATCGTAGATATGTCAAGATGCATGTTTTGATTAGGGGCATGGCCCACCAAATACTCCAAACGAACTCTCCAGTCATGGTTGGGGATTTTCGT
GGAGGATTACCAGATGTGGATATGTGGAAAGAGAATCTAGTGAGAGTCTCTCTGAAGCATTGTTATTTCAAGGAAATTCTTTCTAGTCATTCACCAAGAT
GTCCCAATCTTTCAACTTTATTGCTATGTGATAATGAAGGGTTGCAATTTATTACAGGTTCTTTTTTCACGCAATTGCATGGGCTCAAGGTTCTTGACCT
GTCTCATACAAACATCACAAAATTGCCTAATTCTGTCTCTGAACTGGTGAGTCTCACTGCATTGTTGCTCAAAGAATGCAGAAATTTAAGGCATGTACCA
TCATTAGAAAAGCTCAGAGCACTGAAGAGGTTAGATCTCTCTGGTACTCGGTCACTTGAAATGATGCCTCAAGGCATGCAATGTCTATCCAACCTGAGGT
ATCTGAGGATGAATGGATGTGGTGAAAAGGAGTTTCCTCCTGGGATATTACCAAATCTTTCTCAACTGAAAGTCTTTATATTAGAGGAGATTGATTATGA
CTACTTTCCAGTAACAGTTGAAGGAAAGGAAGTAGGATGCTTGTGGGAGTTGGAAAATTTGGAATGCTATTTTAAGGGTCAATCTGACTTTGTGGAGTAT
CTCAATTCTCGGAATAAGATCCAATCACTAAGAAAATACCATATTTTCGTAGGATCACGGGATAAAGGTTGTGATCGTGAAATAGATGATGGTGAAATGA
AGCTTGATTATGGCATAAGTAAAACAGTTTCGTTGGGTAATTTGCGTAACCAAGGAGATGGAGATTTTCAAGTCATGTTCCCAAAAGACATTCAACAACT
GGTTATTTATGATTGTTCATGTGATGTTTCCTCTCTAATAGAGCATTCAACAGAACTGGAGGTCATCCACATTGAGGATTGCAGTAGCATGGAGAGCTTG
ATTTCATCTTCTTGGTTCTACCCTTCTCCAACACCATTGCCATCTTATAATGGTGTATTTTCTGGTCTTAAAGTGTTCAATTGCTCTGGATGTAGTCGTA
TGAAGGAGTTGTTCCCTCTTGTCTTGCTGCCAAACCTCGTAAACCTAGAAAAGATTACAGTTAGAGATTGTGAGAAGATGAAGGAGATAATAGGTGGAAC
AAGATCAGATGAAAAAGGGGTTATGGGTGAAGAAAGCAACAACAACAGCTTTGGACTCAAATTGCCAAAGTTAAGAGAACTGACATTGAGGGGATTACCA
GAACTGAAAAGCATTTCTAGTGCAAAACTGATTTGCGATTCCCTGGAATTAATTGAAGTATTGTATTGTGAGAAGCTGAAGAGGATGCCAATATGTCTTC
CGTTGCTTGAAAATGGCCAGCCATCTCCTCCCCCTTCTCTTAGAAGAATTGAGATATGTCCAGAAGAATGGTGGGAGTCAGTAGTGGAGTGGGAGCATCC
TAACACTACATATGTCCTTCGTCCCTTTGTAAAGGTTCAGGAGACGATATTGCCCCTTGATTTGGATATTGAATTTCGGGGCCGAAAAAGAAAGGGGATG
GAAATGGTTATACAGAATGATCCATTTTGGGAATATGTTGAGAAGCTGGATGGTGGAAGTTTCAAGTGTACATTTTGTGCCTATAAATTTGCTACTGCTA
CTTCTGTTTCGAGGATCAAATGGCATTTGGCAGGGCGTGGTGTTAAATTATGTGACAAAGTTCCTGAAGAAGTTCAAGATGCAGCCCGTGCAGCAATTGA
TGGTCCTCCAGAAAAGAAACAAAAAGTTATAGCAGGTTCAAGCAATAACGATGGCAATAATGCAATTACAACTTCTGCACAAGAACAAAACTATGAAGGG
AGACATGTAGAAATGGCACAACAAGGAGAAGCTTTTTCCCCTACAGCGCTCAAAGTTTGTTTGGATAGCATCGTTGATAAAGAGATTGAGTTTAGGCATG
ATGCATCTGAGACTATACCGATAACCGAACAAGTGCAGAATCTGAAGAGAGGTAGCTCTCTTGAGAGGCCATCAATTAATCAAGCTGACGGGCCTCGAGG
AGATTCATCCCCACCAAAGGATCTATCGTGTCTTGGCCTTGGAAGTTATCATGATCAACTCTGTTCTCCATCAGTAAAAAACGATGTCATGATGGATGAT
GTGCAGAACATAGTTAGGGAGAAAACAGAGCCAGTAGCTAGTATGTTAGAGCAAAGCAATACAATACTTAATAAATTGGCAGGTGATGATGGAAGGATAC
AGGTGGGAGTTCAGGGCAAGGAACAAGGTGCAGAGGAAGAGTTGATTTGTTCGCATCCAGAAGCGGAAAGTAGCATGGAATACACCTGTGAAGGATTTAT
TCAGCAAATTGACAGAAATGTCTCCCCAGAACGAGCTAGATTAATGGAGAACAGTAGTGGAAGATTAGTGCAAACCGGCACAAGTGCTAGCTCTACAAAG
CTAGTGGGTCAAGCATTTGAACAGAATATGAAGGTGATAAGGTCTTGGTTAATGGATGATGAAGTCTCAACTATTGGCATTTACGGAATGGGGGGAGTTG
GTAAAACAACAATGCTGCAACAAATCTGTAATGAGCTTCTAGGAAGACCAGGCATTTCTCAAGATGTTTGCTCGGTGACTATATCTCAAGACTTTAATAT
TAAAACATTGCAAAATCTCATTGCTAAACGTCTTGATTTAGATATTTCAAGCGAAGACGATGATAAGAGTAAAGCTGTCAAATTGGCAAAAGAACTAGAG
AAGAAACAAAAATGGATTCTCATTTTAGATGATTTGTGGAACTCTTTTGAGCCACAAGAAGTGGGGATTCCTATCTCATTGAAAGGATCCAAGCTCATTA
TGACAACTCGATCAGAAATGGTTTGTCGGCAGATGAATAGCCAAAACAATATAAGAGTGGATCCTCTTTCTGATGAAGAATCTTGGACTTTGTTCATGGA
GAAGCTTGGACAAGACAAACCACTTTCTCCAGAAGTGGAACGAATTGCTGTAGATGTTGCAACGGAATGTGCTGGTTTGCCATTGGGAATTGTTACATTA
GCAGAAAGTTTGAAGGGAGTGAATGACCTATTTGAGTGGAGGATTACATTGAAGAGATTGAAAGAATCAAATTTTTGGCACATGGAAGATCAGATATTCC
AGATATTAAGATTGAGTTATGATTGTTTAGATGATGCAGCACAACAATGCTTCGCATATTGTGCATTATTTGATGAATGTCATAAGATTGAAAGGGAGGA
GTTGATAAAGTCGTTTATCGAGGAGGGGATAATTAAAGAAATGAACAATGGCCACTCAATTCTTGATAGACTAGAAGATGTCTGCTTATTGGAAAGAATT
GATGGTGGTAGTGCTGTTAAGATGCATGACTTGCTCAGGGACATGGCCTTGCACATACTGGATGAGTACTCTCTAATCATGGGCTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.018G145532.1 pacid=42800318 polypeptide=Potri.018G145532.1.p locus=Potri.018G145532 ID=Potri.018G145532.1.v4.1 annot-version=v4.1
MTCKFCGHCFTQGTSSRRIKLHLAGVRGRGVKICKNVPQEVRDAACEAVNDSPPEKKLETAAGSSTNEVANAISASTQEQNIEVTYVEMALQGGPFFTRE
LAWANDLRGSSHDSSRPADDQLCSSVNNDVNMKDVQNKVGVKTEPVLFQVVEQSNAEPVSLAGHAGSIQVGVHGMEQGAEEERICPNLAANGMENPCDGS
SQHSRRLIIDAQDNTNSAFLIKEEDVENNLGRLVQPGAGASSSGGVACKTNKIKGDALLTRKMVGQAFNDHKETIQSLLEHNEVSSIGIYGMGGVGKTTL
VTHIYNQLLERRDTHVYWITGSQDTSINRLQTSLARRIGLDLSSEDEELHRAVALKKELMKKQKWVLILDDLWKAFDLQKLGVPDQVEGCKLILTTRSEK
VCQQMKTQHTIKVQPISEEEAWTLFIERLGDDIALSSEVKRIAVDIVRECAGLPLGIITMARSMRGVDDPYEWTDTLKKLKESKCREMEDEVFQLLRFSY
DQLNDLALQQCLLYCALYPEDHEIEREELIGYLIDEEIIEEMRSRQAAFDEGHMMLDKLEKVCLLERVGYRRYVKMHVLIRGMAHQILQTNSPVMVGDFR
GGLPDVDMWKENLVRVSLKHCYFKEILSSHSPRCPNLSTLLLCDNEGLQFITGSFFTQLHGLKVLDLSHTNITKLPNSVSELVSLTALLLKECRNLRHVP
SLEKLRALKRLDLSGTRSLEMMPQGMQCLSNLRYLRMNGCGEKEFPPGILPNLSQLKVFILEEIDYDYFPVTVEGKEVGCLWELENLECYFKGQSDFVEY
LNSRNKIQSLRKYHIFVGSRDKGCDREIDDGEMKLDYGISKTVSLGNLRNQGDGDFQVMFPKDIQQLVIYDCSCDVSSLIEHSTELEVIHIEDCSSMESL
ISSSWFYPSPTPLPSYNGVFSGLKVFNCSGCSRMKELFPLVLLPNLVNLEKITVRDCEKMKEIIGGTRSDEKGVMGEESNNNSFGLKLPKLRELTLRGLP
ELKSISSAKLICDSLELIEVLYCEKLKRMPICLPLLENGQPSPPPSLRRIEICPEEWWESVVEWEHPNTTYVLRPFVKVQETILPLDLDIEFRGRKRKGM
EMVIQNDPFWEYVEKLDGGSFKCTFCAYKFATATSVSRIKWHLAGRGVKLCDKVPEEVQDAARAAIDGPPEKKQKVIAGSSNNDGNNAITTSAQEQNYEG
RHVEMAQQGEAFSPTALKVCLDSIVDKEIEFRHDASETIPITEQVQNLKRGSSLERPSINQADGPRGDSSPPKDLSCLGLGSYHDQLCSPSVKNDVMMDD
VQNIVREKTEPVASMLEQSNTILNKLAGDDGRIQVGVQGKEQGAEEELICSHPEAESSMEYTCEGFIQQIDRNVSPERARLMENSSGRLVQTGTSASSTK
LVGQAFEQNMKVIRSWLMDDEVSTIGIYGMGGVGKTTMLQQICNELLGRPGISQDVCSVTISQDFNIKTLQNLIAKRLDLDISSEDDDKSKAVKLAKELE
KKQKWILILDDLWNSFEPQEVGIPISLKGSKLIMTTRSEMVCRQMNSQNNIRVDPLSDEESWTLFMEKLGQDKPLSPEVERIAVDVATECAGLPLGIVTL
AESLKGVNDLFEWRITLKRLKESNFWHMEDQIFQILRLSYDCLDDAAQQCFAYCALFDECHKIEREELIKSFIEEGIIKEMNNGHSILDRLEDVCLLERI
DGGSAVKMHDLLRDMALHILDEYSLIMG
|
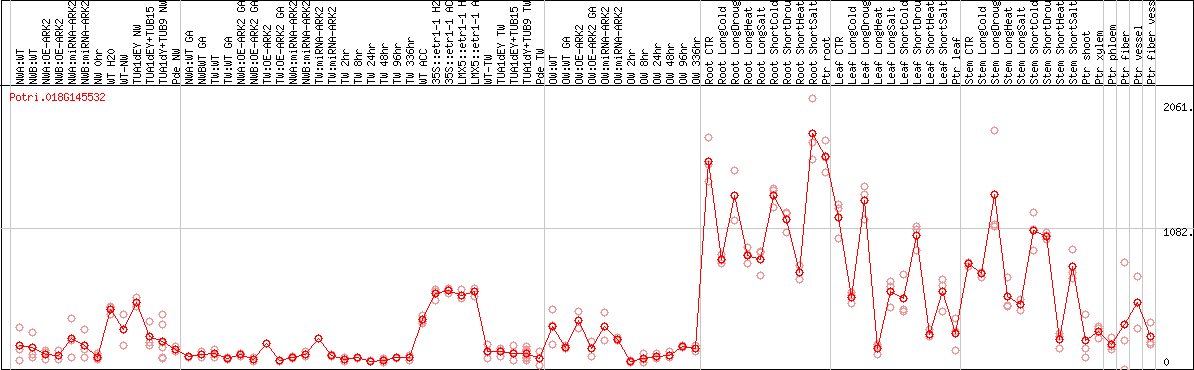
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.018G145532 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
