External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G36930 182 / 3e-52
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
AT4G16990 161 / 2e-45
RLM3
RESISTANCE TO LEPTOSPHAERIA MACULANS 3, disease resistance protein (TIR-NBS class), putative
AT5G17680 159 / 2e-44
disease resistance protein (TIR-NBS-LRR class), putative (.1)
AT5G41540 152 / 6e-42
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT1G27170 152 / 7e-42
transmembrane receptors;ATP binding (.1.2)
AT5G49140 151 / 2e-41
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT5G46450 149 / 5e-41
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT5G58120 149 / 7e-41
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT1G56540 149 / 8e-41
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT3G44630 149 / 1e-40
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2), Disease resistance protein (TIR-NBS-LRR class) family (.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.T005501
406 / 1e-134
AT5G36930 612 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.019G003285
394 / 1e-129
AT5G36930 640 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.019G003942
383 / 7e-126
AT5G36930 591 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.007G143100
254 / 8e-85
AT5G36930 234 / 1e-69
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.003G084333
254 / 2e-82
AT5G36930 347 / 2e-108
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.T002568
252 / 4e-81
AT5G36930 439 / 2e-142
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.019G019710
252 / 5e-81
AT5G36930 445 / 2e-144
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.013G037599
262 / 9e-81
AT5G36930 627 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.T002428
251 / 2e-80
AT5G36930 387 / 2e-122
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10039850
193 / 6e-56
AT5G36930 573 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10018616
190 / 5e-55
AT5G36930 578 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10011104
188 / 3e-54
AT5G17680 574 / 0.0
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Lus10015453
181 / 5e-54
AT1G27170 282 / 5e-84
transmembrane receptors;ATP binding (.1.2)
Lus10014582
172 / 2e-52
AT1G27170 186 / 9e-56
transmembrane receptors;ATP binding (.1.2)
Lus10032101
174 / 1e-51
AT1G27170 270 / 6e-80
transmembrane receptors;ATP binding (.1.2)
Lus10042752
163 / 8e-51
AT4G16990 145 / 5e-41
RESISTANCE TO LEPTOSPHAERIA MACULANS 3, disease resistance protein (TIR-NBS class), putative
Lus10005171
174 / 1e-49
AT1G27170 576 / 0.0
transmembrane receptors;ATP binding (.1.2)
Lus10018309
167 / 2e-49
AT1G27180 245 / 1e-71
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Lus10041060
170 / 4e-48
AT5G17680 641 / 0.0
disease resistance protein (TIR-NBS-LRR class), putative (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0023
P-loop_NTPase
PF00931
NB-ARC
NB-ARC domain
CL0173
STIR
PF13676
TIR_2
TIR domain
Representative CDS sequence
>Potri.019G001314.1 pacid=42773999 polypeptide=Potri.019G001314.1.p locus=Potri.019G001314 ID=Potri.019G001314.1.v4.1 annot-version=v4.1
ATGGCTTCCACGTGCTCTGAGTCTCCGTTCTTTTCTTCGTCTTCATCCTCTAGACATAGATGGAACTATGATGTTTTTCTAAGTTTTAGAGGAAAAGATA
CCCGCAAGAACTTTACAGATCATCTCTATACCGCCTTAATCCAGGCCGGAATTCATACTTTTAGGGATGACAATGAACTTCCTAGAGGAGAAGAAATCTC
TCCACAGCTTGTCAAGGCAATTGAAGGATCAAGGATTTCTATAGTGGTCTTCTCCAAACAATACGCTTCCTCTAGATGGTGTCTTGACGAACTGGTAAAG
ATTGTTGAGTGCAGGCAAAAGATTGACCAGGTTGTTCTTCCTATATTCTATGATACGGAGCCTTCAGATGTCCGAAAACAGACCGGGAGTTATGCAAAAG
CTTTTGATGAACATGAAGAACACTTCAAGGAAGAAATGGAGAAGGTCAACAAGTGGAGAGGAGCTCTTGCAGAAGCGGGAAATCTATCTGGATGGGGTTT
AAACAATGAGGCAAATGGATATGAAGCGGAATTCATCAAAAGGATTGTCAGTGATGTAGCTTGTAAACTGGGAAACAAAACCTTGCACGTAGCCAAGCAC
CCAGTAGGTATTTATTCCAGGGTTCAGGGAATTATTTCTTTGCTAAAAGGTGCAAAACCTGATGTTGGCATCGTGGGAATACATGGGATAGCTGGAATAG
GCAAGACAACAATAGCAAAAGCTGTGTTAACAAACTTTATTTGGATTTGA
AA sequence
>Potri.019G001314.1 pacid=42773999 polypeptide=Potri.019G001314.1.p locus=Potri.019G001314 ID=Potri.019G001314.1.v4.1 annot-version=v4.1
MASTCSESPFFSSSSSSRHRWNYDVFLSFRGKDTRKNFTDHLYTALIQAGIHTFRDDNELPRGEEISPQLVKAIEGSRISIVVFSKQYASSRWCLDELVK
IVECRQKIDQVVLPIFYDTEPSDVRKQTGSYAKAFDEHEEHFKEEMEKVNKWRGALAEAGNLSGWGLNNEANGYEAEFIKRIVSDVACKLGNKTLHVAKH
PVGIYSRVQGIISLLKGAKPDVGIVGIHGIAGIGKTTIAKAVLTNFIWI
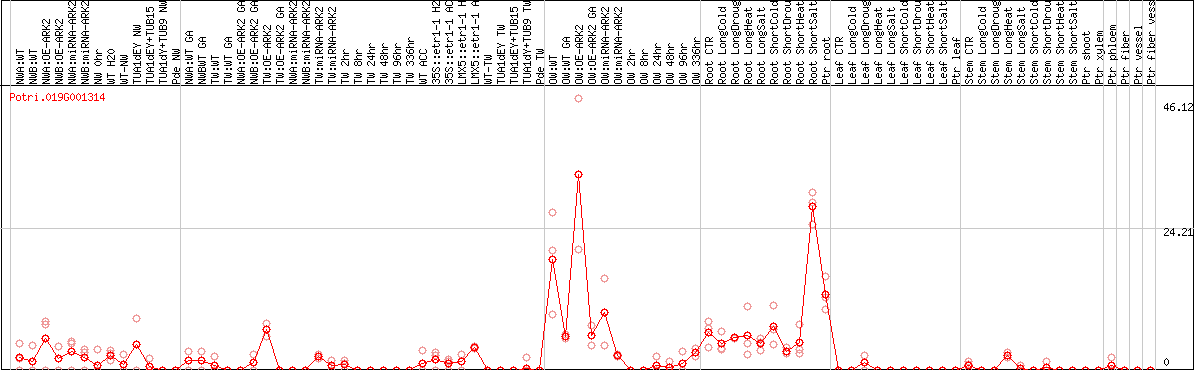
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.019G001314 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.