Potri.019G003285 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.019G003285.1 pacid=42774306 polypeptide=Potri.019G003285.1.p locus=Potri.019G003285 ID=Potri.019G003285.1.v4.1 annot-version=v4.1
ATGGCTTCCACGGACTCTGAGTCTCCGTTCTCTTCTTCGTCTTCATCCTCTGGACATAGATGGAACTATGATGTTTTTCTAAGTTTTAGAGGAGAAGATA
CCCGCAAGAACTTTACAGATCATCTCTATACCGCCTTAATCCAGGCCGGAATTCATACTTTTAGGGATGATAATGAACTTCCTAGAGGAGAAGAAATCTC
TCCACAACTTCTCAAGGCAATCGAAGGATCAAGGATTTCTATAGTGGTCTTCTCCAAACACTACGCTTCCTCTAGATGGTGTCTTGACGAACTTGTAAAG
ATTCTTGAGTGCAGGCAAAACATTGACCAGGTTGTTCTTCCTATATTCTATGATACGGAGCCTTCAGATGTCCGAAAACAGACCGGGAGTTATGCAAAAG
CTTTTGATGAACATGAAGAACGCTTCAAGGAAGAAATGGAGAAGGTCAACAAGTGGAGAGGAGCTCTTGCAGAAGCGGGAAATCTATCTGGATGGGGTTT
ACACAATGAGGCAAATGGATATGAAGCGGAATTCATCAAAAGGATTGTCAGTGATGTAGCTTGTAAACTGGGAAACAAAACCTTGCACGTAGCCAAGCAC
CCAGTAGGTATTTATTCCAGAGTTCAGGGTATTATTTCTTTGCTAAAAGGTGCAAAACCGGATGTTGGCATCGTGGGAATACATGGGATAGCTGGAATAG
GAAAGACAACAATAGCAAAAGCTGTGTTTAACAAACTTTATTTTGGATTTGAGGGAAGCAGTTTTCTTTCGGACGTCAAGGAAATATCAGATAAGCCGAA
TGGTCTGGTTGAATTACAAGAGCGCCTTCTTCATGATATTCTGAAACCAAGAGTTTGGAAGGTCAGTAATGTTTATGAAGGCATGAATTTGATCAAAGAA
CGACTTCATCGTAAGAAAATTCTTGTTGTTTTTGACGATGTGGACAAAAGGGAACAACTGGAGGCATTGATGGGAGAGAGATGCTGGTTTGGTGCCGGAA
GTATAATAATTGTTGTAACTAAGAATAAGCATCTGCTAACAGAAGTGGGGGTAGATGGAATGTATCATGCTAAAGAATTGGATCGAGATCAGTCTCTTGA
GCTTTTCAGTTTGCATGCTTTTAGGGAAACCCATCCAGCAAAGGATTATGAGGAGCTTTCAGGAAAGGTAGTTGACTACTGCAAAGGACTTCCTTTGGCT
CTACAAATTTTGGGTTCTCATTTATCTATAAGAGACAAAGCTGGATGGGAAATCGATATTGCCCATTGGCGAAATATTCCGCATGATGATATTCAAGGAA
AGCTTAGAGTAAGTTTTGATGCACTGAATGTGGATACAAGTGAGATATTCCTTGATATTGCGTGCTATTTTGTTGGTGGAGACAAAGAATATGTAGCAGA
TATAGTAGGTGCTCGTTACGATTGCCATCCAGAAGTAGCTTTCAGAACTCTCATTGGAAGGTCTCTCATAACAATCGACACTTGGAATAGCTTATGGATG
CATGATACATTACGAAAGATGGGAAGGGAGATTATTCGTCAAAGGTCGCGTAATCATCCTGGAAATTGCAGCAGAATTGTACTTCCCAAAGACGCATACA
ATGTACTCTCCAAGGAACTGGGAACAGATGCTGTGGAGGGTCTGGCACTAGATGTGCAAGAATCTTTTAGCACAAAATCGTTTACAAAAATGAGACGTTT
AAAATTACTCCAAATCAAAGGAGCAAATCTTGTGGGAAGCTACAGTCTTCTCCCCAAAGAGTTGATATGGCTTTGTTGGTTTGGATGCCCTTTGAAATCT
TTGCCGTCTGATTTTCACTTAAACGACCTTGTTATACTTGATATGCAAGAAAGTAACGTACGAAAACTTTGGAAGGGGACTAAGATTCTTAACAAGCTGA
AAATCCTCAATCTCAGCTATTCCAAGTACCTTGATGAAACGCCGAACTTTCGAGAACTCTCTTGTTTGGAGAGACTAATACTTACTGGATGCACGAGTTT
GGTTAAGGTACATCAATCTATTGGAAATTTGAAGAGCCTTGTTTTGTTGAACTTGCATTACTGTGACAGCCTAAAGACTCTTCCAGAAAGCATGGGTAAC
TTAAAATCTCTTCAAACTCTGAACGTTACTCAGTGCAGACAACTTGAGAAATTGCCAGAGAGTTTGGGTGACATAGAATCCTTAACAGAATTGTTTACAA
AGGGTACTGCGATTAAGCAACTTCCTACTTCAGCTAGATATTTGAAGAAGCTGACAAAGTTATCATTTGGTGGATACAACAAAGTTTTCTACTCACCTGA
TCTACCTTCTAAATCTCGGTTTTCAAGATTTTCTTTGTGGCTTTCACCACGAAACTGTTCTAGCTCCAACGCTATGCTACCAGCTTTTTTCAATAGTTTC
AGCTCATTGAAAGAACTAAATCTAAGTTATGCTGGTTTGTCTGAAGCTACAAGTTCCATTGATCTTGGGAGTTTGTCCTTTCTTGAAGATTTGGATTTGT
CTGGAAACAAATTCTTCAATCTGCCTTCTGGCATTAGCCTCCTTCCTAAGCTTCAATGTTTGAGGGTTGAGAAATGTTCGAATCTGCTATCAATCCCAGA
GCTTCCATCAAGTGTCCTATTCTTGTCTATCAATGACTGCACATCCATTGAAAGAGTAAGTGCACCACTTCAGCACGAAAGACTACCATTACTGAATGTA
AAAGGGTGCCGAAATTTAATAGAGATTCAGGGCATGGAATGCGCGGGCAATAACTGGTCCATTTTGAACTTAAATGGCTGCAGCAATTTATCTGAAAACT
ATAAGATGAGTCTTATTCAGGGATTATGCAAAGGTAAACATTATGACATTTGCCTTGCTGGTGGTGAGATACCAGAATGGTTCAGCCATCGTGGAGAAGG
ATCTGCATTATCATTTATTTTACCCTCAGTGTCAGTTCCAGTTCCAGATGGTAATAAACTCCAAGCGCTCCTTTTTTGGGTTGTCTCGGCCTCCACCAAT
GAAGCCACTCTAGCAACATCACTTCCTCGGTTTGATAAGTGTGTTGCTACTTTCAAAAATAAGAGTAATGGTATTGAATTGTTTGAGACGATGGCGGCAG
TTACGTTTGACAGAACTATCACGAAGCAGTCTTGGATACAGCGTATACCGTTGATTGGGTTAGAGGAATCGCTGCAAGGTGTAGAGGAATTGGAAGTGAA
TGTCAAAATAAGCGTATATGATGTTCGAAAATTACAGCGTATACCGTTGATCGGGTCAGAGGAATCGCTGCAAGGTGTAGAGGAATTGGAACTGAATGTC
AAAATAAGCTTCTATGATGTTCGAAAATGTTGGGTAGAAAAATGCGGGGTACATTTGATAATGGAAAAGAATAAAGCAGATTCAGATCAGGAGATTGATA
TTCATGCTCCAGGCTCTGATGATCAGCCGTTGGAAAGTAGTTTGATAAGAGAGTTGCAAAAATGGAAAATCACCAGTTGCAGTAAATTTGGATAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.019G003285.1 pacid=42774306 polypeptide=Potri.019G003285.1.p locus=Potri.019G003285 ID=Potri.019G003285.1.v4.1 annot-version=v4.1
MASTDSESPFSSSSSSSGHRWNYDVFLSFRGEDTRKNFTDHLYTALIQAGIHTFRDDNELPRGEEISPQLLKAIEGSRISIVVFSKHYASSRWCLDELVK
ILECRQNIDQVVLPIFYDTEPSDVRKQTGSYAKAFDEHEERFKEEMEKVNKWRGALAEAGNLSGWGLHNEANGYEAEFIKRIVSDVACKLGNKTLHVAKH
PVGIYSRVQGIISLLKGAKPDVGIVGIHGIAGIGKTTIAKAVFNKLYFGFEGSSFLSDVKEISDKPNGLVELQERLLHDILKPRVWKVSNVYEGMNLIKE
RLHRKKILVVFDDVDKREQLEALMGERCWFGAGSIIIVVTKNKHLLTEVGVDGMYHAKELDRDQSLELFSLHAFRETHPAKDYEELSGKVVDYCKGLPLA
LQILGSHLSIRDKAGWEIDIAHWRNIPHDDIQGKLRVSFDALNVDTSEIFLDIACYFVGGDKEYVADIVGARYDCHPEVAFRTLIGRSLITIDTWNSLWM
HDTLRKMGREIIRQRSRNHPGNCSRIVLPKDAYNVLSKELGTDAVEGLALDVQESFSTKSFTKMRRLKLLQIKGANLVGSYSLLPKELIWLCWFGCPLKS
LPSDFHLNDLVILDMQESNVRKLWKGTKILNKLKILNLSYSKYLDETPNFRELSCLERLILTGCTSLVKVHQSIGNLKSLVLLNLHYCDSLKTLPESMGN
LKSLQTLNVTQCRQLEKLPESLGDIESLTELFTKGTAIKQLPTSARYLKKLTKLSFGGYNKVFYSPDLPSKSRFSRFSLWLSPRNCSSSNAMLPAFFNSF
SSLKELNLSYAGLSEATSSIDLGSLSFLEDLDLSGNKFFNLPSGISLLPKLQCLRVEKCSNLLSIPELPSSVLFLSINDCTSIERVSAPLQHERLPLLNV
KGCRNLIEIQGMECAGNNWSILNLNGCSNLSENYKMSLIQGLCKGKHYDICLAGGEIPEWFSHRGEGSALSFILPSVSVPVPDGNKLQALLFWVVSASTN
EATLATSLPRFDKCVATFKNKSNGIELFETMAAVTFDRTITKQSWIQRIPLIGLEESLQGVEELEVNVKISVYDVRKLQRIPLIGSEESLQGVEELELNV
KISFYDVRKCWVEKCGVHLIMEKNKADSDQEIDIHAPGSDDQPLESSLIRELQKWKITSCSKFG
|
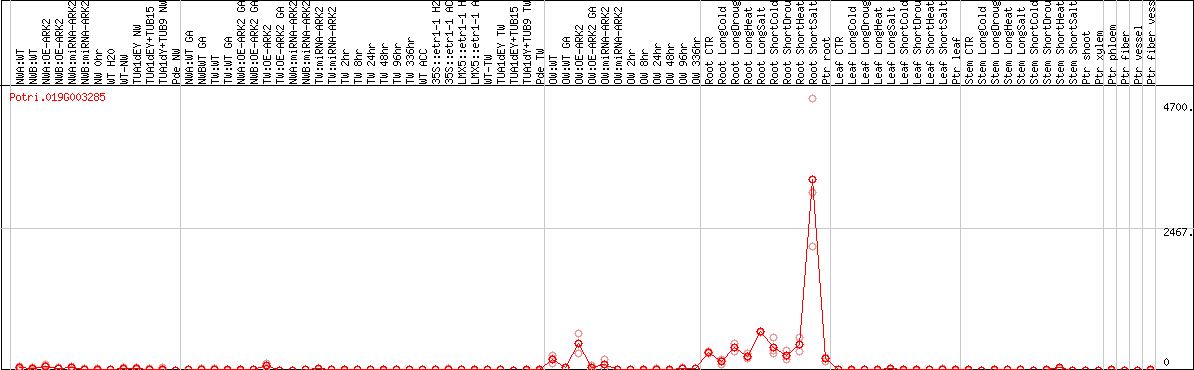
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.019G003285 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
