Potri.019G014322 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.019G014322.1 pacid=42773883 polypeptide=Potri.019G014322.1.p locus=Potri.019G014322 ID=Potri.019G014322.1.v4.1 annot-version=v4.1
ATGGAAGGAGTCCCACCGTCTTGTAATGAGGATGATGTCAACTTGACATCGAATCAGTTCGATTTTCAATCACCAGAAGATTCATTCAATATTTCTGAGT
TTCTCTCGACGCCACCGGAGTTTCCAGGCAACCAAAGCTTCATGTTATGCAATGATCATGTTGGTCAGAACATGGTGGTGGATAACAATGTGATCAACGA
TAGTATTCAACAAACAACTGATCGGTTTCCACCTTCATTTCCTCAAACCATGACAACAATGGGAGATTTAGATGGAACATCATCGTCATGGATCAACCAA
AATGAACCAAATTGGCCTCAAGCTCCACCAAGGTGTCAGCCTAATCCTAATTTATTTGATCCTCAATGTGCCAATCCTTTGAGGGACCAGCAATGGACTC
ATTATCAGTTACCATTGAGCAATCAAGTGCCTGCGATGTTACCTGTACCTCAACCTCCAATGAATCAACGGCATCAGAACTCCTTTATTCATCAACCTCG
GCCGGGACATGGCTATATGACACCGAACGCAACCACTTCTCAACATGGTGGCTTCAATCAATCTGGACCTTCTTCTCCTGCTCCCTGGTCACAGATGGAT
AACCTACAAGTCCGAGGATTGCAAAATCAAATTGCTAGGCCAAATGCCTCAAATACTCAACAAGTCGAAACGGTTGGATCAAAGTATCCATTTCGGATTC
ATGTTGAATATAGGGCTGATGGTAGCATGAAGTGTAAGTTTTGTCAACGTACATGGGCCAATAAAACTTCCATTTCAAGGTTCAAATGGCATTTATCAGG
AGAGGAAGGGCGTGGTGTTGCCATTTGTCGTGGGGTGACTAAAGAAGTTAAAGAAGCAGCCTTTCTAGCTATATGTGGTGGCCACAAAAGACAAAAAATC
ACAGCAAGTTCAGTCAATGTTAATGACTGTGGAATTCCAACTTGTCTACAAGAACTAAACATTGAGAATGAAAATACGGGAGGTGTTGGAAGGGTACAAA
GAGAAGTTCAGGTTGTAGAACCCGGAGTAGTGGAAGAGAGGATTTCTTCACATGCAATAGCAGGAAACGACGTAGTAAGCATGACCGGAATGAGAGCACA
AGAAGATAGAGTTTCCGAAGGGGCACTCGAGAGTAGACTGAGGACAGAACCAGTGGATCGATCGTTGGAACAAAGCAATGCAGTACTTGGCAATATGGCA
GGGGGTGCTGGAAGGATACAAGTGGGAGTTCAGGTCATGGAACAAGGTCCCGGAGAAGAGAGAATTCAGTCGCATCTACAAGCGGAAAATGGCATGGAAA
ACACTTGTGAAGGATCTTTTCAGCATGATGCATTTGAGACTGTACCAAGAACAGAGCAAGTGCAGCTTCTGGAGCCTCGAGGAGATTCATCCCAATTTTG
TCGTGACATTGGAAGATGTTATGATCAACCTTGTGCTCCATCAGTAAACAATGATGCAAATAGGCATGATGCGCAGGACATGGTTAGAGTGAGGACACAA
CCAGTGCAAGAAGAGGAGGATGTGGAGAATAGTGGAAGATCAATAGTGCAGGCTGGCGCAGGAGCTAGATCTTCTGAAAGTCTGAAGTACAACAAGACTA
GAGGAGTTCCATTACCTACTAGCTCTACAAACCCAGTGGGTCAAGAATTTGAAGAGAATTCGAAGGTGATATGGTCTTTGTTAATGGATGGGGAAGTCTT
AAGCATGGGTATTTATGGGATGGGGGGAGTTGGTAAATCCACAATACTTCAACATATCTACAATGAACTTCTACAAAAACCAGATATTTGTGATCATGTT
TGGTGGGTGACCGTGTCTCAAGATTTCAGCATTAATAGATTGCAGAATCTTATTGCTAAACATCTTGATCTAGACCTTTCAAGAGAAGATGATGATCTGT
ATAGAGCTGCCAAATTGTCAGAAGAACTAATGAAGAAACAAAAATGGATTCTCATTTTAGATGATTTGTGGAACAATTTTGAGCTACACAAAGTGGGAAT
TCCTGAGAAGTTAGAAGGATGCAAGCTGATTATAACAACTCGATCAGAAATGATTTGTCATCAGATGGCTTGCCAGCACAAAATCAAAGTGAAGCCACTT
TCTAATGGAGAAGCTTGGACTTTGTTCATGGAGAAACTTGGACGTGATGTAGCACTTTCACCAGAAGTGGAAGGAATTGCGAAAGTTGTTGTAAGGGAAT
GTGCTGGTTTGCCATTGGGAATTATTACAGTGGCAGGAAGCTTGAGGGGAGTGGATGATCTACATGAGTGGAGGAATACATTGAATAAACTGAAAGAATC
AGAATTTAGGGACATGGATGAGAAGGTATTCAAGTTATTGAGGTTTAGTTATGATCAGTTAGGTGATTTAGCACTACAACAATGTCTCTTGTACTGTGCA
TTATTTCCTGAAGATGATGATATTGAAAGGGAGGAGTTGATAGGTTATTTGATCGACGAGGGAATAATTAAAGGAAAAAGGAGAAGGGAAGATGCATTTG
ACGAGGGCCACACGATGCTTAATAAACTTGAATATGTGTGCCTATTAGAAAGAGCTCAAATGATGTTTGGTTGTAGATGTGTCAAGATGCATGATTTGAT
TAGGGACATGGCCATCAAAATACTGCTAGAGAACTCTCAAGGCATGGTCAAAGCAGGTGCACAATTAAAAGAGTTGCCAGATACAGAGGAGTGGACGGAG
AAGCTGACGAGAGTTTCATTAATGCAAAACGAGATCAAAGAAATTCCTTCCAGCCATTCACCTAGGTGTCCCTTTCTATCAACTCTATTGCTATGTCGTA
ATCGTTGGTTGGGATTTATTTCGGATTCATTTTTCAAGCAATTGCATGGGCTCAAGGTCCTCGATCTGTTTTACACGAGTATTGAAAATTTGCCAGACTC
TGTCTCTGATTTGGTGAGCCTCACAGCATTATTGCTCAATGATTGTAAGAAATTAAGACATGTTCCATCATTAGAGAAGCTCACGGCACTGAAGAGGTTG
AATCTCTCTCGTACTGCACTTGAAAAGATGCCGCAAGGAATGGAATGCCTAACCAACCTGACGTATCTTAGAATGAATGGATGTGGTGAAAAGGAGTTTC
CTAGGGGGATATTACCAAAACTCTCTCACCTGCAAGTCCTTGTACTAGAAGACTTTTTTGATGGAAGTTATGCTCCGATAACAGTTGAAGGAAAGGAAGT
AGGATCCTTGAGAAATTTGGAAAGTTTGGAATGCCATTTCGAAGGTTTTTCTGACTTCGTGGAGTATGTCAGATCTTGGGATGGGATCCTATCATTAAGC
ACATACAGAATTTTAGTAGGAATGGTGGGTGGATTTATAGGTCAACACATTGAATATTTTCCAAGTAAAACAGTTGGGTTGGGTAATTTGAGTATCAACG
GAGATAGAGATTTTCAGGTCAAGTTCTTAAATGGCATTCAAGGATTGATTTGTGGATGCATCGATGCAAGAAGTTTATGTGATGTTTTCTCATTAGAGAA
TGCAACTGAACTGAAGCTCATCAGCATTTGGAAATGCCATAACATGGAGAAGCTGTTCCCGCTTGTGTTGCTGCCAAACCTCGTAAACCTGGAAAGGATT
GAAGTGATGTTCTGTGAGAAAATGGAGGAGATAATAGGAACAACAGATGAAGAAAGCAACACCTCCAATTCCATCAAGGAAGTCATTCTCCCAAAGTTAA
GATTTCTGAAATTGATTGGGTTACCAGAACTGAAAAGCATTTGCAGTGCAAAACTGATTTGCAATTCTATTAAACATATTATTGTAAGGTGGTGTGAGAA
GTTGAAGAGGATACCAATTTGTCTTCCGTTGCTTGAAAATGGCCAGCCATCTCCTCCCCCTTCTCTTGAAAATATCTATTCAAGTCCAGAAGAATGGTGG
GAGACAGTAGTGGAGTGGGAGCATCCTAATGCAAAGGATGTCCTTCGTCCCTTTGTATGCTACTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.019G014322.1 pacid=42773883 polypeptide=Potri.019G014322.1.p locus=Potri.019G014322 ID=Potri.019G014322.1.v4.1 annot-version=v4.1
MEGVPPSCNEDDVNLTSNQFDFQSPEDSFNISEFLSTPPEFPGNQSFMLCNDHVGQNMVVDNNVINDSIQQTTDRFPPSFPQTMTTMGDLDGTSSSWINQ
NEPNWPQAPPRCQPNPNLFDPQCANPLRDQQWTHYQLPLSNQVPAMLPVPQPPMNQRHQNSFIHQPRPGHGYMTPNATTSQHGGFNQSGPSSPAPWSQMD
NLQVRGLQNQIARPNASNTQQVETVGSKYPFRIHVEYRADGSMKCKFCQRTWANKTSISRFKWHLSGEEGRGVAICRGVTKEVKEAAFLAICGGHKRQKI
TASSVNVNDCGIPTCLQELNIENENTGGVGRVQREVQVVEPGVVEERISSHAIAGNDVVSMTGMRAQEDRVSEGALESRLRTEPVDRSLEQSNAVLGNMA
GGAGRIQVGVQVMEQGPGEERIQSHLQAENGMENTCEGSFQHDAFETVPRTEQVQLLEPRGDSSQFCRDIGRCYDQPCAPSVNNDANRHDAQDMVRVRTQ
PVQEEEDVENSGRSIVQAGAGARSSESLKYNKTRGVPLPTSSTNPVGQEFEENSKVIWSLLMDGEVLSMGIYGMGGVGKSTILQHIYNELLQKPDICDHV
WWVTVSQDFSINRLQNLIAKHLDLDLSREDDDLYRAAKLSEELMKKQKWILILDDLWNNFELHKVGIPEKLEGCKLIITTRSEMICHQMACQHKIKVKPL
SNGEAWTLFMEKLGRDVALSPEVEGIAKVVVRECAGLPLGIITVAGSLRGVDDLHEWRNTLNKLKESEFRDMDEKVFKLLRFSYDQLGDLALQQCLLYCA
LFPEDDDIEREELIGYLIDEGIIKGKRRREDAFDEGHTMLNKLEYVCLLERAQMMFGCRCVKMHDLIRDMAIKILLENSQGMVKAGAQLKELPDTEEWTE
KLTRVSLMQNEIKEIPSSHSPRCPFLSTLLLCRNRWLGFISDSFFKQLHGLKVLDLFYTSIENLPDSVSDLVSLTALLLNDCKKLRHVPSLEKLTALKRL
NLSRTALEKMPQGMECLTNLTYLRMNGCGEKEFPRGILPKLSHLQVLVLEDFFDGSYAPITVEGKEVGSLRNLESLECHFEGFSDFVEYVRSWDGILSLS
TYRILVGMVGGFIGQHIEYFPSKTVGLGNLSINGDRDFQVKFLNGIQGLICGCIDARSLCDVFSLENATELKLISIWKCHNMEKLFPLVLLPNLVNLERI
EVMFCEKMEEIIGTTDEESNTSNSIKEVILPKLRFLKLIGLPELKSICSAKLICNSIKHIIVRWCEKLKRIPICLPLLENGQPSPPPSLENIYSSPEEWW
ETVVEWEHPNAKDVLRPFVCY
|
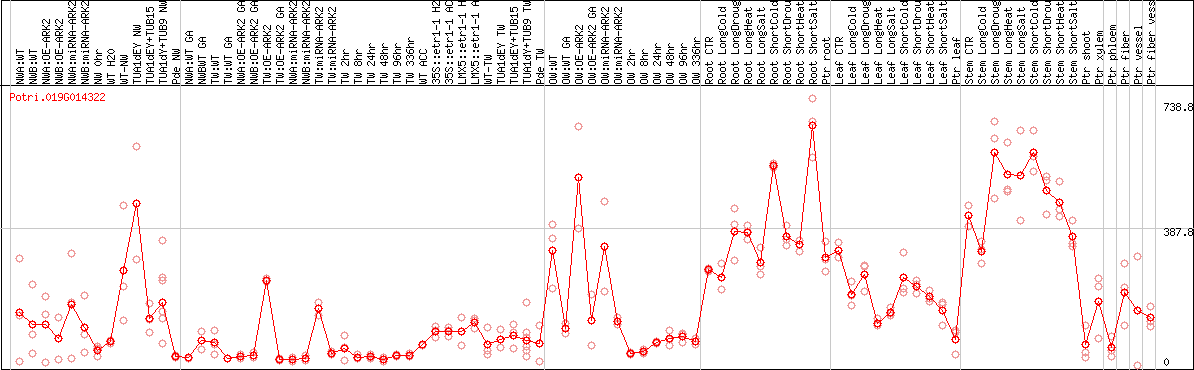
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.019G014322 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
