Potri.019G014328 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.019G014328.1 pacid=42773421 polypeptide=Potri.019G014328.1.p locus=Potri.019G014328 ID=Potri.019G014328.1.v4.1 annot-version=v4.1
ATGTCATATAATATAATCACGCACTTGAAGAACATCTTGAATCAGATGGAAGGGTTCCCACCATCTTCTGCGGAGGATGATGTCATGATCAACTTGACAT
CGAATCAGTTCGATATTCAATCACCAGAAGATTCATTCAATATATCTGAGCTTCTCTCGTCGCCACTGGAGGATCTAGCCAACCAAAGCTTCATATCATG
CAATGATCAAGTTGAACAGAACTTCGTGGTGGATAACAATGTGATCAACGATAGTATTCAGCAAACAACTCAGTTTCCACCTTCATCTTCTCAAGCTATA
ATGGAAGGGGTCACAGCATCTTCTATTGTGGATGATGTCAACTTGAGATCGAATGAGTCAGATGGAAGGCCTCAGCTTCTCTTAAGCAGGAAGGAGGTTT
CAGGCAATGATCAAGGTGGACAGAACAACGTGATCAATGATAGTATTCAACAAACAACTCAGTTTCCACTTTCATTTCCTCAAACCATGACAACAATGGG
AGGTTTCGATGGAACATCATCATCACGGACCAACCAAAATGAACCAAATTGGCCTCAAACTCCTTCAAGGGACCAACAATGGACTAATTATCAGTTACCA
ATGAGCAATCCAGTGCCTGCCATGCTACCTGGACCTCAACCTCCAATGAATCAATGGCATCATAATTCCTTTATTCATCAACCTCAGCCGGGACATGGCT
ATATGACACCGAATGCAACCACTTCACAACATGGTGGCTTCAATCAATTTGAACCTTCTTTTCCTGCTCTCCGGAGTCAATATTTTTCTAATCAAGGACA
AGTCGATCAAGCTGCTGTAATTCCTAACAAAGACAATGTGCAACAAGAGAACTTCGTGCCTAGGTCACAGATGGATAACCTCCACGTCCGAGGATTGCAA
AATCAAACTGCTAGGCCAAATGCCTCAAATCCTGGCCTTGACACTTCCTTGCAACGTCAAATTACGGGTTTAAATACTCAACAAGTCGAAATGGTTGGAT
CAAATGATCCATTTCGGAATTATGTTGAAGAAAGGGCTGATGGTGGTAGAATGAAGTGTAAGTTTTGTCCACATACATATGCCATTAAAACTTCCATTTC
AAGGATCAAATGGCATTTATCAGGAGAAGAAGGGCATGGTGTTGCCATTTGTCGTTGGGTGCCTAAAGAAGTTCAAGAAGCAGCCTGGAAAGCCATGTGT
GGTGGCAACAAAAGACATAAAATCACAGCAAGTTCAATCAATGTTAATGACTGTGGAATTTCAACTTGTCCACAAGAACAAAACATTGAGGTTAACATGG
GAGGAGGTATTAGAAGGGTACAAGGAGAAGTTCAGGTTGTAGAACCGGGAGTAGGGGAAGAGAGGATTTCTTCACAAGCAATAGCAGGAAACGACGTAGT
AAGCATGACCGGAATGAGAGCAACAGAAGATGGAGTTTCCGAAGGGGCACTGGAGAGCAGACCGAGGACAGAACCAGTGGATCGAGCGTTGGAACAAAGC
AATGCAGTACTTGGCAATTTGGCAGGGGGTGCTGGAAGGATACAAGTGGGAGTTCAGGGCATGGAACAAGGTGCTGGAGAAGAGAGAATTCAGTTGCATT
TACAAGCGGAAAATGGTATGGAAAACACTCCTGAAGGATGTTTTCGGCATGATGCATCTGAGACTATACCAAGAACAGAGCAAGTGCAGCTTCAAGAGGC
TCGAGGAGATTCATCCCAATTTTGTCTTGACACTGGAAGATATTATGATCAACTCTGTGCACCATCAATAAGCAAAGATGTCCTTATGTATGATGTGCAG
AACATGGTTAGAGTGAGGACAGAACCAGTGGAGGAGGAGGGTGTGGAGAATAGTGGAAGATTAGTGCTGCCTGGCGCAGGAGCTGGATCTTCTAGAAGTC
TTAAATACAACACAAGTGAGACTAGAGGAGTTCCATTACCTACTAGCTCTACAAAACCAGCGGGTCAAACATTTGAAGAGAATACGAATGTGATATGGTC
TTTGTTAATGGATGATGAAGTCTCAGTCATTGGTATTTATGGAATGGGAGGAGTTGGTAAAACAAAAATACTGCAACATATTTATAATGAGCTTCTACAA
ATACCAGATATTTGTGATCATGTTTGGTGGGTGACTGTGTCTCAAGATTTCAGCATTAATAGACTGCAAAATCTTATTGCTGAACATCTTGATCTAGATC
TTTCAAGAAAAAATGTTGAACTGCATAGAGCTGCTAAATTATCAGAAGAACTAACGAAGAAACAAAAATGGATTCTCATTTTAGATGATTTGTGGAATAA
TTTTGAGCTCCAAGAAGTGGGAATTCCTATCCCGTTGAAAGGATGCAAGTTAATTTTGACAGCTCGATCAGAAATGGTTTGTCGTCGGATGGCTTGCCAA
CACAAAATCAAAGTGAAGCCACTTTCTGAGGGAGAAGCTTGGACTTTGTTCATGGAAAAACTAAGATGTGAGAAAGCATTTTCACCAAAACTGGAAGGAA
TTGCGAAAGCTATTGCAAGGGAATGTGCTGGTTTACCATTGGGAATTATTACAGTGGCAGGAAGCTTGATGGGAGTGGATGATCTGCATGAGTGGAGCAA
TACATTGAAGGAATTGAGTGAATCAAAATTTAGGGACATGGATGAGAACGTATTCAAGTTATTGAGGTGTAGTTATGATCGGTTAGGTGATTTAGCACTA
CAGCAATGTCTCTTGTACTGTGCATTATTTCCTGAAGATGGCTGCATTGAGAGGGAGGAGTTGATAGGTTATTTGATCGATGAGGGAATAATTAAAGGAA
TGAGGAGCTGGAAAGATGCATTTGACAAGGGCCACACGATGCTTAATAGACTCGAATATGTCTGTCTATTGGAAGGTGATAAATTCAAATATGGTGGTGG
TACATTTGTCAAGATGCATGACTTGATTAGGGACATGGCTATCCAAATACAGCTAGAGAACTCTCAAGTCATGGTTAAAGCTGGTGCGCAATTAAAAGTG
TTGCCAGATGCTGAGGAGTGGACAGAGAATCTCACAAGAGTCTCACTAATGCAAAACCATATCAAAGAAATTCCTTCCAGTCATTCACCAAGGTGTCCAA
ATCTATCAACTTTATTTCTAAATGATAATGACTGGTTGGGATTTATTGCGGATTCATTTTTCAAGCAATTGCATGAGCTCAAGGTCCTCGATCTGTCTGG
CACAAGTATTAAAAATTTGCCGGACTCTGTCTCTGATTTGGTGAATCTCACTACATTATTGATCAAATATTGTGAGAACTTAAGACATGTTCCATCATTA
AAGAAGCTCAGAGAATTGAAGAGGTTAGATCTCTCGTTGACTATGCTTGAAAAGATGCCAGAAGGAATGGAATGTCTATCCAAACTGAGGTATCTTAGAA
TGAATGGATGTGGTGAAAAGGAGTTTCCTAGTAGGATATTACAAAAACTCTCTCACCTGCAAGTCTTTGTACTAGAGGAAGCGTCCATTGATGGAAGATG
TGCTCCGATATCAGTTAAAGTAGAGGATGTAGTATCCTTGAGAAATTTGGAAACTTTGGAATGCTATTTCGAAGGCTTCTCTGACTTCGTGGAGTATCTC
AGATCTCGGGATGGGATTCAATCATTAAGCACATATAAAATTTTAGTAGGAATGGTGGATGAAAATTATTGGGCAGACATTGATAATTTTCCAAGTAAAA
TAGTTGGGTTGGGTAATTTGAGTATCAACGAAGATGGAGATTTTCAGGTTAAGTTCTTAAATGGCATTCAAGGACTGGTTTGTCAATTCATCGATGCAAG
AAGTTTATGTGATGTTTTGTCATTAGAGAATGCAACTGAACTGGAGCTCATCAACATTTACGATTGCGATATCATGGAGAGCTTGGTTTCATCTTCTTGG
TTATGCTCTGCTCCACTACCATTGCCATCATATAATGGTATGTTTTCTGGTCTTAAAGAGCTTTATTGTGGTGGATGTAACAGTATGAAGAAGTTGTTCC
CACTTGTGCTGCTGCCAAACCTCGTAAACCTGGAAAGGATTCTTGTTCGTGATTGTGAGAAAATGGGGGAGATAATAGGAACAACAGATGAAGAAAGCAG
CAGCTCCAATTCCATCACGGAAGTCATTCTCCCAAAGTTAAGAACTCTGGAATTGTTTATGTTACCAGAACTGAAAAGTATTTGCAGTGCAAAACTGATT
TGCAATTCTCTTGAAGTTATTACTGTAATGGGTTGTAAAAAGCTGAAGAGGATGCCAATTTGTCTTCCGTTGCTTGAAAATGGCCAGCCATCTCCTCCTC
ATTCTCTTAGAAGAATGCGTATAAAACCAAAAAAATGGTGGAATACAGTAGTGGAGTGGGAGCATCCTAACGCGAAGGATGTCCTTCGTCCCTTTGTTAA
GTTTCGGTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.019G014328.1 pacid=42773421 polypeptide=Potri.019G014328.1.p locus=Potri.019G014328 ID=Potri.019G014328.1.v4.1 annot-version=v4.1
MSYNIITHLKNILNQMEGFPPSSAEDDVMINLTSNQFDIQSPEDSFNISELLSSPLEDLANQSFISCNDQVEQNFVVDNNVINDSIQQTTQFPPSSSQAI
MEGVTASSIVDDVNLRSNESDGRPQLLLSRKEVSGNDQGGQNNVINDSIQQTTQFPLSFPQTMTTMGGFDGTSSSRTNQNEPNWPQTPSRDQQWTNYQLP
MSNPVPAMLPGPQPPMNQWHHNSFIHQPQPGHGYMTPNATTSQHGGFNQFEPSFPALRSQYFSNQGQVDQAAVIPNKDNVQQENFVPRSQMDNLHVRGLQ
NQTARPNASNPGLDTSLQRQITGLNTQQVEMVGSNDPFRNYVEERADGGRMKCKFCPHTYAIKTSISRIKWHLSGEEGHGVAICRWVPKEVQEAAWKAMC
GGNKRHKITASSINVNDCGISTCPQEQNIEVNMGGGIRRVQGEVQVVEPGVGEERISSQAIAGNDVVSMTGMRATEDGVSEGALESRPRTEPVDRALEQS
NAVLGNLAGGAGRIQVGVQGMEQGAGEERIQLHLQAENGMENTPEGCFRHDASETIPRTEQVQLQEARGDSSQFCLDTGRYYDQLCAPSISKDVLMYDVQ
NMVRVRTEPVEEEGVENSGRLVLPGAGAGSSRSLKYNTSETRGVPLPTSSTKPAGQTFEENTNVIWSLLMDDEVSVIGIYGMGGVGKTKILQHIYNELLQ
IPDICDHVWWVTVSQDFSINRLQNLIAEHLDLDLSRKNVELHRAAKLSEELTKKQKWILILDDLWNNFELQEVGIPIPLKGCKLILTARSEMVCRRMACQ
HKIKVKPLSEGEAWTLFMEKLRCEKAFSPKLEGIAKAIARECAGLPLGIITVAGSLMGVDDLHEWSNTLKELSESKFRDMDENVFKLLRCSYDRLGDLAL
QQCLLYCALFPEDGCIEREELIGYLIDEGIIKGMRSWKDAFDKGHTMLNRLEYVCLLEGDKFKYGGGTFVKMHDLIRDMAIQIQLENSQVMVKAGAQLKV
LPDAEEWTENLTRVSLMQNHIKEIPSSHSPRCPNLSTLFLNDNDWLGFIADSFFKQLHELKVLDLSGTSIKNLPDSVSDLVNLTTLLIKYCENLRHVPSL
KKLRELKRLDLSLTMLEKMPEGMECLSKLRYLRMNGCGEKEFPSRILQKLSHLQVFVLEEASIDGRCAPISVKVEDVVSLRNLETLECYFEGFSDFVEYL
RSRDGIQSLSTYKILVGMVDENYWADIDNFPSKIVGLGNLSINEDGDFQVKFLNGIQGLVCQFIDARSLCDVLSLENATELELINIYDCDIMESLVSSSW
LCSAPLPLPSYNGMFSGLKELYCGGCNSMKKLFPLVLLPNLVNLERILVRDCEKMGEIIGTTDEESSSSNSITEVILPKLRTLELFMLPELKSICSAKLI
CNSLEVITVMGCKKLKRMPICLPLLENGQPSPPHSLRRMRIKPKKWWNTVVEWEHPNAKDVLRPFVKFR
|
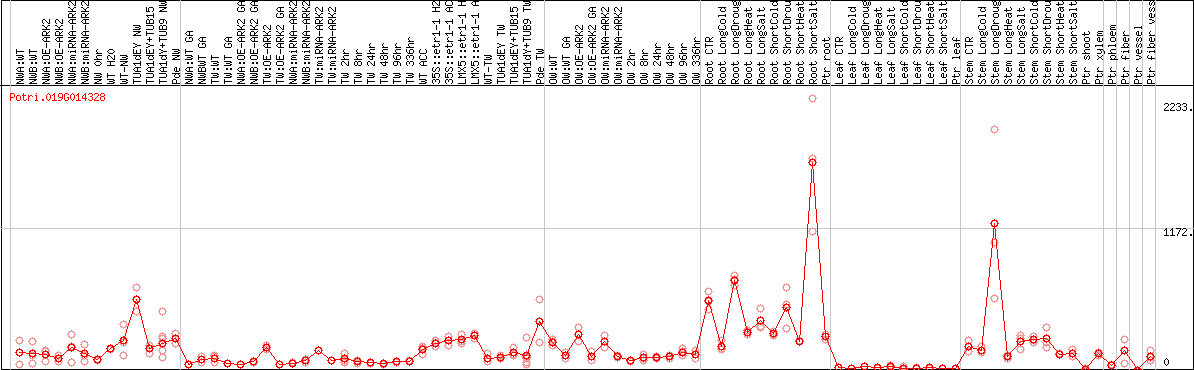
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.019G014328 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
