Potri.019G097100 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.019G097100.3 pacid=42773157 polypeptide=Potri.019G097100.3.p locus=Potri.019G097100 ID=Potri.019G097100.3.v4.1 annot-version=v4.1
ATGGCAATGTTTTTCTTGTTTCTCTCTATCTTTTTGTTTGTTTATATCTGGAGATTCATCATACGAAGCCGTCATATTTCTCCATCCCCATCTACTCCTT
CCACCTTAACCACCGCCCAACCACAAGTAATTAAATATGATGTCTTCCTTAGTTTCAGGGGAGAAGACACTCGTTTTGATTTTACCAGCCACCTCTATGC
TGCTTTGAAACGGAAACAAATCCTAACTTTCATAGATTATCAGCTTGTGAGAGGAGATGAGATTTCGGCATCACTTCTGAGAACAGTTGAAGAGGCCAAG
CTTTCTGTGATTGTTTTTTCTGAAAACTATGCATCTTCCAAATGGTGCTTGGAGGAGCTTGCAAAGATTTTTGAACGTAGAAAAAATAATGGACAGATAG
TTATTCCGGTATTCTACCAGGTGGATCCATCCCATGTAAGAAATCGAACAGGTAGCTTTGGGGATGCATTTGCTAGACTTATAAAGAAGAAAGCTTTGAC
GATGGACAAAGAGCAGAACTTCAGAGACGCTTTGACGGATGCAGCCAATCTATCTGGATGGAGCTTAGGGAAGTCTGAGCTGGAGTCTGAATTTATTGAG
AAAATCGTTGAAGATGTTTTGAACAAATTGCATGACATGTCTTCAAGTCACACTACGGGTTTATTTGGAATTGATGTTCGCGTTAAAAAAGTTGAGTCTT
TGTTAAATATGGAATCTCAAGATGTTCTCATTGTAGGAATATGGGGAATGGGTGGTATCGGTAAGACAACGATTGCTGAAGCTGTGTGCAACAAGGTTCG
TTCTCGATTTGAAGGAATTTTTTTTGCAAACTTTAGGCAAGAATTAGAACCAGGTTCCATGGCTGATTTGCAAAGAAGCTTTCTTTCACAGCTGCTTGGC
CAAGAAATTTTGAACATGGGCTCCTTGAGTTTTCGAGATTCTTTTGTGAGGGAGAGACTTCGTCGTAAAAAGGTTTTTATTCTTCTGGATGATGTGGATG
ATTTAATGCCTTTGGAAGAATGGAAAGATTTGCTTGATGGACGACACAGTTCATTTGGTTTGGGCAGTAAAGTTCTCATAACAAGTAGGGACAAGCAAGT
GCTTAATAATATAGTAGATGAAACTTACGAGGTTGAGAGGTTGAACTATGAAGAAGCTCTTCAACTCTTTAGCTCAAAAGCCTTGAAGAATTGCATCCCT
ACAATTGATCATAGAGACTCGATAAAAAGGATTGCAAGTCATGTACAAGGCAATCCGTTGGCTCTTATAGTTTTGGGTTCCTCTCTCTATGGTAAAAGCC
CAGAAGAATGGTACAGTGCATTGAATAAACTAGCTCTGAACTCTCGAATCGAAAATGCATTGAGAATTAGTTACAATGGGTTGGATCAAGAACAACGATC
CATATTTCTTGATATAGCACATTTCTTTAGAGGATTTGAGCAAAACCAAGCGACAAGAATATTAGATGGCTTTTATGGTCGGCCTGTGATTTTCGATATA
AGCACGCTCATTGATAAGTGTCTCATAACTACTTCTCGTAACATGCTAGAAATACATGATTTACTACAAGAAATGGCATTTAGCATTGTTCGTGCAGAAT
CTAAGTTTCCTGGCAAACGTAGCAGGTTGTGTCATTTTACTAACGTCGTTCATGTATTGGAGGAAAATAAGGGAACTGAAGAAATTGAAGGCATATCTTT
GGACATGTCCAGGTTATCGAGACAGATACACTTGAAATCTGATGCCTTCGCAACGATGAATGGTCTTAGATTTCTCAAATTCTATTTCGGTCATTTCTCC
GAAGATAACAAAAATAAAATGCACCTTCCTCCTACTGGCCTCAAATATCTTTCTAATAAGCTGAGATATTTGCACTGGGATGGATTCCCTTCGAAATCAT
TGCCACAAGTTTTTTGTGCTGAACTCCTTGTTGAGCTTAACTTACGAAGGAGCAAGGTTGAAAAACTTTGGGAAGGAGTACAGGATGTTGGAAATTTACG
AAAATTTGTACTATCTTTCTCTCCATATTTGATGGAATTGCCAGATCTATCAAAGGCCAAAAATTTAGTGTGCTTATACCTTGTGGACTGTTCAAGTTTA
ACTGAGGTTCCGTCATCTCTTCAATCTCTTGACAAGCTGGAAGAACTTGATCTCTATTTTTGCTATAATCTTAGAAGTTTTCCAATGCTTGATTCAAAGG
TTCTCAGAGTACTTGCTATAAGTCGGTGCCTAGATATGACCAAGTGTCCAACTATTTCACAAAATATGAAAAGCTTATATTTAGAGGAAACTTCAATCAA
AGAAGTTCCACGATCAATTACTAGCAAGTTGGAAAGTCTTGGTCTGCATGGTTGCTCAAAGATCACCAAGTTTCCAGAAATCTCAGGGGATATAAAAACA
TTATATCTAAGTGGAACTGCTATTAAAGAGGTGCCCTCATCAATCCAGTTTCTCACAAGACTCCGCGTATTGGATATGAGTGGTTGCTCAAAACTTGAGA
GCTTCCCAAAAATTTTAGTGCCTATGAAATCTTTGGTGGATCTTAACTTGAGTAAAACAGGCATTAAAGAGATACCCTCATCATTTAAGCATATGATATC
TTTGAGAAGTTTAGGGTTAGATGGAACGCCTATTGAAGAGCTACCCTTATCAATCAAAGATATGGTATGTTTGAGATATCTGGCTTTGCATGGAACACCT
ATTAAAGCATTACCTGAGCTTCCACCTTCACTGAGGTCTCTAACAACACATGACTGTGCATCTCTAGAAACCATGATATCAACCATCAATATAGGCAGAT
TATGGGATGGATTGAATTTTGCAAATTGCTTCAAATTGGATCAGAAACCACTGATAACAGCAATGCATTTAAAAATTCAGTCCGGAGATAAGATCCCATA
TGACAGAATGCAAATGGTTCTACCGGGGAGTGAAATTCCGGAATGGTTCGGTGACAAAGGAATTGGATCTTCACTCACCATACAGTTGCCCTCAAATTGT
CGTCAACTCAAGGGAATTGCTTTCTGCCTTGTCTTTCTACTCCCTCTTCCCTCCCATGAGATGCTTTATGAATTCGATGATCATCCTGAAGTTCGCGTAT
ATTTTGATTGCCAAGTTAAGAGTAAAAACGGTGAGCATGATGGTGATGATGAAGAAGTCTTCGTTTCCAAGAAAAGCTATAGTATATTCAATTTCTTGAA
GACATGTGACTCAGATCACATGTTTCTACACTACGAGCTCGAATTAGTAAATCATTTCCGTAAATATTCTGGCAATGAAGTTACATTTAAATTCTACCAT
GAAGTGGTCAACGGGAGTACGAATGTAGGTCATGAGATTCGAAAGCCTTGCAAGTTGAAAAGCTGTGGTGTATATCTGCACTTTGATGAAAATCTACAGG
CCGACACTTTGTTAAGGATCTTCCTTAACAAACAAAAGTTTAGAAGAAAATTGAGGGAGAAATAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.019G097100.3 pacid=42773157 polypeptide=Potri.019G097100.3.p locus=Potri.019G097100 ID=Potri.019G097100.3.v4.1 annot-version=v4.1
MAMFFLFLSIFLFVYIWRFIIRSRHISPSPSTPSTLTTAQPQVIKYDVFLSFRGEDTRFDFTSHLYAALKRKQILTFIDYQLVRGDEISASLLRTVEEAK
LSVIVFSENYASSKWCLEELAKIFERRKNNGQIVIPVFYQVDPSHVRNRTGSFGDAFARLIKKKALTMDKEQNFRDALTDAANLSGWSLGKSELESEFIE
KIVEDVLNKLHDMSSSHTTGLFGIDVRVKKVESLLNMESQDVLIVGIWGMGGIGKTTIAEAVCNKVRSRFEGIFFANFRQELEPGSMADLQRSFLSQLLG
QEILNMGSLSFRDSFVRERLRRKKVFILLDDVDDLMPLEEWKDLLDGRHSSFGLGSKVLITSRDKQVLNNIVDETYEVERLNYEEALQLFSSKALKNCIP
TIDHRDSIKRIASHVQGNPLALIVLGSSLYGKSPEEWYSALNKLALNSRIENALRISYNGLDQEQRSIFLDIAHFFRGFEQNQATRILDGFYGRPVIFDI
STLIDKCLITTSRNMLEIHDLLQEMAFSIVRAESKFPGKRSRLCHFTNVVHVLEENKGTEEIEGISLDMSRLSRQIHLKSDAFATMNGLRFLKFYFGHFS
EDNKNKMHLPPTGLKYLSNKLRYLHWDGFPSKSLPQVFCAELLVELNLRRSKVEKLWEGVQDVGNLRKFVLSFSPYLMELPDLSKAKNLVCLYLVDCSSL
TEVPSSLQSLDKLEELDLYFCYNLRSFPMLDSKVLRVLAISRCLDMTKCPTISQNMKSLYLEETSIKEVPRSITSKLESLGLHGCSKITKFPEISGDIKT
LYLSGTAIKEVPSSIQFLTRLRVLDMSGCSKLESFPKILVPMKSLVDLNLSKTGIKEIPSSFKHMISLRSLGLDGTPIEELPLSIKDMVCLRYLALHGTP
IKALPELPPSLRSLTTHDCASLETMISTINIGRLWDGLNFANCFKLDQKPLITAMHLKIQSGDKIPYDRMQMVLPGSEIPEWFGDKGIGSSLTIQLPSNC
RQLKGIAFCLVFLLPLPSHEMLYEFDDHPEVRVYFDCQVKSKNGEHDGDDEEVFVSKKSYSIFNFLKTCDSDHMFLHYELELVNHFRKYSGNEVTFKFYH
EVVNGSTNVGHEIRKPCKLKSCGVYLHFDENLQADTLLRIFLNKQKFRRKLREK
|
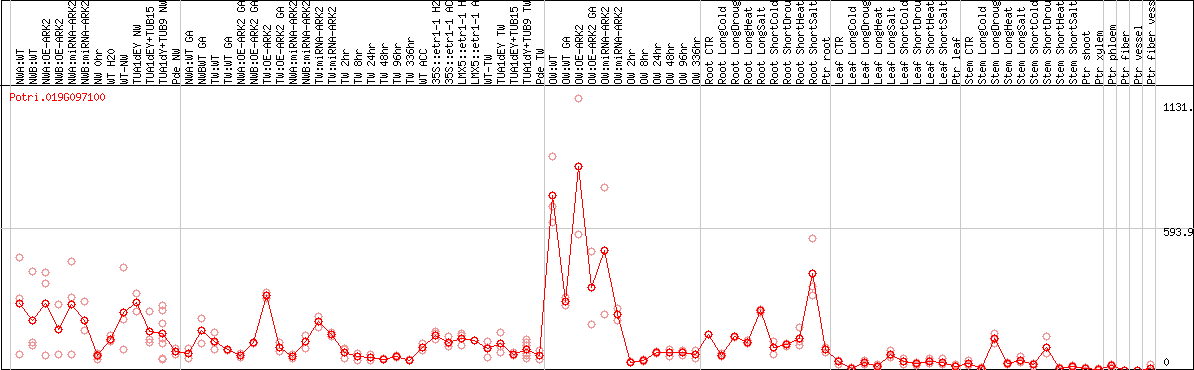
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.019G097100 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
