Potri.019G098900 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.019G098900.1 pacid=42774603 polypeptide=Potri.019G098900.1.p locus=Potri.019G098900 ID=Potri.019G098900.1.v4.1 annot-version=v4.1
ATGGCGTTGTTTTTCTTGTTTCTCTCTATCTTTTTGTTTGTTTTTATCTGGAGATTCATCATACGAAACTGTCATATATCTCCATCGCCATCTACTCCTT
CCACCTTAACCACCGCCCAGCCACAAGTAATTAAACATGATGTCTTCCTTAGTTTCAGGGGAGAAGACACCCGTGGTGGTTTTACCAGCCACCTCTATGC
TGCTTTGGACCGGAAACAAATCCGAGCTTTCATAGATTATCAGCTTCGGAGAGGAGATGAGATTTCGGCATCACTTCTGAGAACAATTGAAGAGGCCAAG
CTTTCTGTGATTGTTTTTTCTGAAAACTATGCATCTTCCAAATGGTGCTTGGAGGAGCTTGCAAAGATTATTGAGCGTAGAAGAAATAATGGACAGATAG
TTATTCCAGTATTCTACAAGGTGGATCCATCCCATGTAAGAAATCAAACACGAAGCTTTGGAGATGCATTGGCTAGATTGATAAAGAAGAAAGCTTTGAC
GATGGACAAAGAGCAGAGCTTCAGAGACGCTTTGACGGCTGCAGCCAATCTATCTGGATGGAGCTTAGGGAATTCTGAGCTGGAGTTTGAATTTATTAAG
AATATCGTTGGAGATGTTTTGGAAAAATTGCATGCCATGTCTTCAAGTCACACTATGGCGGGTCTACTTGGAATTGATGTTCATGTTAGCAAAGTTGAGT
CTTTGTTAAATATAGAATCTCCAGATGTTCTCATTGTAGGGATATGGGGAATGGGTGGTATCGGTAAGACAACGATTGCTGAAGCTGTATGCAACAAGGT
TCATTCTCAATTTGAAAGAATCTTTTTTGCAAACTGTAGGCAACAATCTGATTTGCCAAGAAGATTTCTTAAACGGCTGCTTGGCCAAGAAACTCTGAAC
ACTATGGGCTCCTTGAGTTTTCTAGATTCTTTTGTGAGGGATAGACTTCGTCGAATTAAGGTTTTCATTGTTCTGGATGACGTGGATGATTTAATGCGTT
TGGATGAATGGAGAGATTTGCTTGATGGACGAAACAATTCATTTGGTTCGGGCAGTAAAGTTCTCATAACAAGTAGGAACAAGCAACTGCTTAAGAATGT
AGTTGATGAGACATACGAGGTTGAGGGGTTGAACTATGCAGACGCTATTCAACTCTTTAGCTCAAAAGCCTTGAAGAATTGCATCCCCACAATTGATCAA
AGACACTTGATAATAAAGAATGTAAGGCATGTACAAGGCAATCCGTTGGCTCTTAAAGTTTTGGGTTCCTCTCTCTATGATAAAAGCATCGAAGAATGGC
GCAGTGCATTGAAGAAACTAGCTCTGGACCCTCAAATCGAAAGGGCGTTGAGAATTAGTTACGATGGGTTGGATTTAGAACAAAAACCCATATTTCTTGA
CATAGCACATTTCTTCAAAGGACGGATGCAAGGCGAAGCAACAGGAATATTAGATTGCTTATATGGTCAGTCTGTGAATTTTGATATAAGCACGCTTATT
GATAAGTGTCTCATAAGTACTGCAAAGGATTACTTTCATCGTGACAAGCTAGAAATGCATGATTTACTACAAGAAATGGCATTTAACATTGTTCGTGCAG
AATCTGATTTTCCTGGCGAACGTAGCAGGTTAAGTCATCCTCCTGATGTCGTTCAATTATTGGAGGAAAATAAGGGTACTCAACAAATTAAAGGCATATC
TTTGGACATGTCCATGTTATCGAGACAGATACACTTGAAATCTGATGCATTCGCAATGATGGATGGTCTTCGATTTCTCAATATCTATTTCAGTCGTTAC
TCCAAAGAAGATAAAATATTGCACCTTCCTCCTACTGGCCTTGAATATCTTCCTAATGAGCTGAGATATTTTTTATGGTCTAGATTCCCTTTGAAATCCT
TGCCGCCATCTTTTCGTGCTGAACACCTTGTCGAGCTTCACCTAAGGAAAAGCAAGCTTGTAAAACTTTGGACAGGAGTAAAGGATGTTGGGAATTTAAG
AAGAATTGACCTATCTGACTCTCCATATTTGACGGAATTGCCAGATCTATCAATGGCCAAAAATTTAGTGTCCTTAGATCTTACGGACTGTCCAAGTTTA
ATCGAGGTTCCGTCATCTCTTCAATATCTTGACAAGCTGGAAAAAATTTATCTCTTTCGTTGCTATAATCTTAGAAGTTTTCCAATGCTTGATTCAAAGG
TTCTCAGATTCCTTCTTATAAGTCGGTGCCTGGATGTGACCACGTGTCCAACGATTTCACAAAATATGGAATGGTTATGGTTGGAACAAACTTCAATCAA
AGAAGTTCCACAATCAGTTACTGGCAAGTTGGAACGTCTTTGTCTAAGTGGTTGCCCAGAGATCACCAAGTTTCCAGAGATTTCAGGGGATATAGAAATA
TTAGATCTAAGGGGAACTGCTATTAAAGAGGTGCCCTCATCAATCCAATTTCTCACAAGACTCGAAGTGTTGGATATGAGTGGTTGCTCAAAACTTGAGA
GCCTCCCAGAAATCACAGTGCCTATGGAATCTTTGCATAGCCTTAAGTTGAGTAAAACAGGCATTAAAGAGATACCCTCATCATTAATTAAGCATATGAT
ATCTTTGACGTTTCTAAATTTAGATGGAACGCCTATTAAAGCGTTACCTGAGCTTCCCCCTTCGCTTCGTTATCTAACAACACATGACTGTGCATCACTA
GAAACCGTGACATCAAGCATCAATATCGGTAGATTAGAGTTAGGATTGGATTTTACAAATTGCTTCAAATTGGATCAGAAACCACTGGTAGCAGCAATGC
ATTTGAAAATTCAGTCCGGAGAGGAGATCCCAGATGGCGGTATTCAAATGGTACTACCGGGGAGTGAAATTCCAGAATGGTTCGGTGACAAAGGAATTGG
ATCTTCACTCACCATGCAGTTGCCGTCCAATTGTCATCAGCTCAAGGGAATTGCTTTCTGCCTTGTCTTTCTACTCCCTCTTCCCTCCCATGACATGCCT
TATGAAGTCGATGATGATATTGATGTTAATTTATATCTTGATTACCATGTTAAGAGTAAAAACGGTGAGCATGATGGTGATGATGAAGTCGTCTTAGCTT
CCGGGGAAAGATGTCATCTAACATCTAAAATGAAGACATGTGACTCGGATCACATGGTTCTACATTACATGGCTCTACGTTACGAGCTCGAATTGGTAAA
TCGTTTACGTAAATATTCCGGCAATGAAGTTACATTTAAATTCTACCACCATGAAGTGGTCAACATGGCGAGGAAGGTAGGTAATGAGATCCAAAGGCCT
TTCAAGTTGAAAAGCTGTGGGGTGTATCTGCACTTTGGTGAAAATCTACCGGCGGACACGGACGGGTACAGATATTTACCCGTTAACAAACAACGGTTCA
GAAGAAAATAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.019G098900.1 pacid=42774603 polypeptide=Potri.019G098900.1.p locus=Potri.019G098900 ID=Potri.019G098900.1.v4.1 annot-version=v4.1
MALFFLFLSIFLFVFIWRFIIRNCHISPSPSTPSTLTTAQPQVIKHDVFLSFRGEDTRGGFTSHLYAALDRKQIRAFIDYQLRRGDEISASLLRTIEEAK
LSVIVFSENYASSKWCLEELAKIIERRRNNGQIVIPVFYKVDPSHVRNQTRSFGDALARLIKKKALTMDKEQSFRDALTAAANLSGWSLGNSELEFEFIK
NIVGDVLEKLHAMSSSHTMAGLLGIDVHVSKVESLLNIESPDVLIVGIWGMGGIGKTTIAEAVCNKVHSQFERIFFANCRQQSDLPRRFLKRLLGQETLN
TMGSLSFLDSFVRDRLRRIKVFIVLDDVDDLMRLDEWRDLLDGRNNSFGSGSKVLITSRNKQLLKNVVDETYEVEGLNYADAIQLFSSKALKNCIPTIDQ
RHLIIKNVRHVQGNPLALKVLGSSLYDKSIEEWRSALKKLALDPQIERALRISYDGLDLEQKPIFLDIAHFFKGRMQGEATGILDCLYGQSVNFDISTLI
DKCLISTAKDYFHRDKLEMHDLLQEMAFNIVRAESDFPGERSRLSHPPDVVQLLEENKGTQQIKGISLDMSMLSRQIHLKSDAFAMMDGLRFLNIYFSRY
SKEDKILHLPPTGLEYLPNELRYFLWSRFPLKSLPPSFRAEHLVELHLRKSKLVKLWTGVKDVGNLRRIDLSDSPYLTELPDLSMAKNLVSLDLTDCPSL
IEVPSSLQYLDKLEKIYLFRCYNLRSFPMLDSKVLRFLLISRCLDVTTCPTISQNMEWLWLEQTSIKEVPQSVTGKLERLCLSGCPEITKFPEISGDIEI
LDLRGTAIKEVPSSIQFLTRLEVLDMSGCSKLESLPEITVPMESLHSLKLSKTGIKEIPSSLIKHMISLTFLNLDGTPIKALPELPPSLRYLTTHDCASL
ETVTSSINIGRLELGLDFTNCFKLDQKPLVAAMHLKIQSGEEIPDGGIQMVLPGSEIPEWFGDKGIGSSLTMQLPSNCHQLKGIAFCLVFLLPLPSHDMP
YEVDDDIDVNLYLDYHVKSKNGEHDGDDEVVLASGERCHLTSKMKTCDSDHMVLHYMALRYELELVNRLRKYSGNEVTFKFYHHEVVNMARKVGNEIQRP
FKLKSCGVYLHFGENLPADTDGYRYLPVNKQRFRRK
|
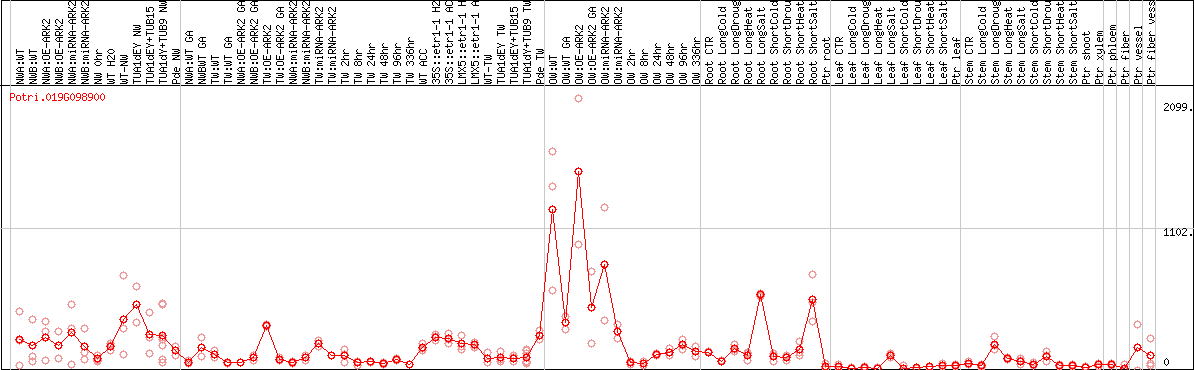
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.019G098900 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
